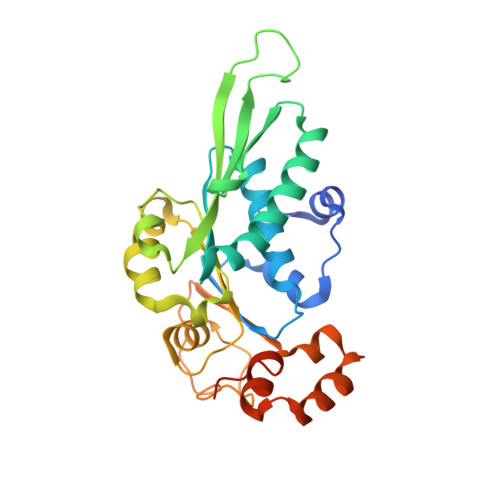
Structure and mechanism of the ATPase that powers viral genome packaging.
Hilbert, B.J., Hayes, J.A., Stone, N.P., Duffy, C.M., Sankaran, B., Kelch, B.A.(2015) Proc Natl Acad Sci U S A 112: E3792-E3799
- PubMed: 26150523 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1506951112
- Primary Citation Related Structures:
4ZNI, 4ZNJ, 4ZNK, 4ZNL - PubMed Abstract:
Many viruses package their genomes into procapsids using an ATPase machine that is among the most powerful known biological motors. However, how this motor couples ATP hydrolysis to DNA translocation is still unknown. Here, we introduce a model system with unique properties for studying motor structure and mechanism. We describe crystal structures of the packaging motor ATPase domain that exhibit nucleotide-dependent conformational changes involving a large rotation of an entire subdomain. We also identify the arginine finger residue that catalyzes ATP hydrolysis in a neighboring motor subunit, illustrating that previous models for motor structure need revision. Our findings allow us to derive a structural model for the motor ring, which we validate using small-angle X-ray scattering and comparisons with previously published data. We illustrate the model's predictive power by identifying the motor's DNA-binding and assembly motifs. Finally, we integrate our results to propose a mechanistic model for DNA translocation by this molecular machine.
- Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Worcester, MA 01605;
Organizational Affiliation: