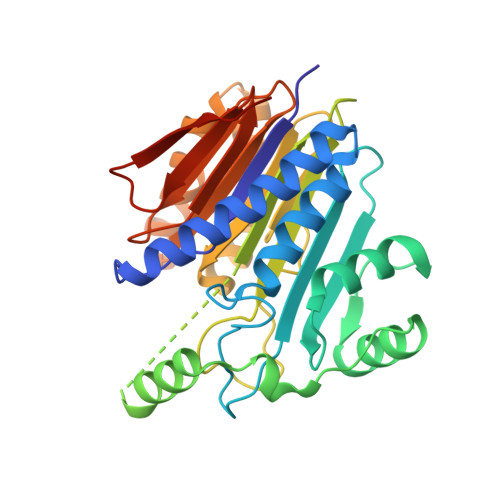
Intramolecular Cleavage of the hASRGL1 Homodimer Occurs in Two Stages.
Li, W., Irani, S., Crutchfield, A., Hodge, K., Matthews, W., Patel, P., Zhang, Y.J., Stone, E.(2016) Biochemistry 55: 960-969
- PubMed: 26780688 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.5b01157
- Primary Citation Related Structures:
4ZM9 - PubMed Abstract:
The human asparaginase-like protein 1 (hASRGL1) is a member of the N-terminal nucleophile (Ntn) family that hydrolyzes l-asparagine and isoaspartyl-dipeptides. The nascent protein folds into an αβ-βα sandwich fold homodimer that cleaves its own peptide backbone at the G167-T168 bond, resulting in the active form of the enzyme. However, biophysical studies of hASRGL1 are difficult because of the curious fact that intramolecular cleavage of the G167-T168 peptide bond reaches only ≤50% completion. We capitalized upon our previous observation that intramolecular processing increases thermostability and developed a differential scanning fluorimetry assay that allowed direct detection of distinct processing intermediates for the first time. A kinetic analysis of these intermediates revealed that cleavage of one subunit of the hASRGL1 subunit drastically reduces the processing rate of the adjacent monomer, and a mutagenesis study showed that stabilization of the dimer interface plays a critical role in this process. We also report a comprehensive analysis of conserved active site residues and delineate their relative roles in autoprocessing and substrate hydrolysis. In addition to glycine, which was previously reported to selectively accelerate hASRGL1 cleavage, we identified several novel small molecule activators that also promote intramolecular processing. The structure-activity analysis supports the hypothesis that multiple negatively charged small molecules interact within the active site of hASRGL1 to act as a base in promoting cleavage. Overall, our investigation provides a mechanistic understanding of the maturation process of this Ntn hydrolase family member.
- Department of Molecular Biosciences, ‡Department of Chemical Engineering, and §Institute of Cellular and Molecular Biology, University of Texas , Austin, Texas 78712, United States.
Organizational Affiliation: