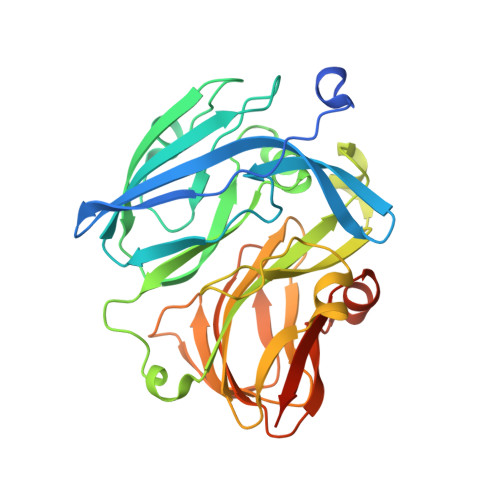
Structural insights into the function of a thermostable copper-containing nitrite reductase
Fukuda, Y., Tse, K.M., Lintuluoto, M., Fukunishi, Y., Mizohata, E., Matsumura, H., Takami, H., Nojiri, M., Inoue, T.(2014) J Biochem 155: 123-135
- PubMed: 24293549 Search on PubMed
- DOI: https://doi.org/10.1093/jb/mvt107
- Primary Citation Related Structures:
4ZK8 - PubMed Abstract:
Copper-containing nitrite reductase (CuNIR) catalyzes the reduction of nitrite (NO(-)2) to nitric oxide (NO) during denitrification. We determined the crystal structures of CuNIR from thermophilic gram-positive bacterium, Geobacillus thermodenitrificans (GtNIR) in chloride- and formate-bound forms of wild type at 1.15 Å resolution and the nitrite-bound form of the C135A mutant at 1.90 Å resolution. The structure of C135A with nitrite displays a unique η(1)-O coordination mode of nitrite at the catalytic copper site (T2Cu), which has never been observed at the T2Cu site in known wild-type CuNIRs, because the mobility of two residues essential to catalytic activity, Asp98 and His244, are sterically restricted in GtNIR by Phe109 on a characteristic loop structure that is found above Asp98 and by an unusually short CH-O hydrogen bond observed between His244 and water, respectively. A detailed comparison of the WT structure with the nitrite-bound C135A structure implies the replacement of hydrogen-bond networks around His244 and predicts the flow path of protons consumed by nitrite reduction. On the basis of these observations, the reaction mechanism of GtNIR through the η(1)-O coordination manner is proposed.
- Department of Applied Chemistry, Graduate School of Engineering, Osaka University, 2-1 Yamadaoka, Suita, Osaka 565-0871, Japan; Department of Chemistry, Graduate School of Science, Osaka University, 1-1 Machikaneyama, Toyonaka, Osaka 560-0043, Japan; Department of Applied Chemistry, School of Engineering, Osaka University, 2-1 Yamadaoka, Suita, Osaka 565-0871, Japan; Faculty of Life and Environmental Sciences, Department of Environmental Information, Kyoto Prefectural University, Shimogamo-Hanki-cho, Sakyou, Kyoto 606-8522, Japan; Molecular-Recognition Structure Analysis Team, Molecular Profiling Research Center for Drug Discovery (molprof) National Institute of Advanced Industrial Science and Technology (AIST) 2-3-26 Aomi, Koto-ku, Tokyo 135-0064, Japan; Microbial Genome Research Group, Japan Agency of Marine-Earth Science and Technology, 2-15 Natsushima, Yokosuka, Kanagawa 237-0061, Japan; and RIKEN SPring-8 Center, 1-1-1 Kouto, Sayo, Hyogo 679-5148, Japan.
Organizational Affiliation: