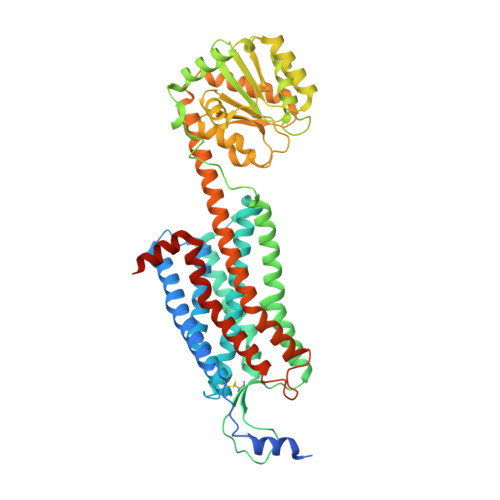
Structure and ligand-binding mechanism of the human OX1 and OX2 orexin receptors.
Yin, J., Babaoglu, K., Brautigam, C.A., Clark, L., Shao, Z., Scheuermann, T.H., Harrell, C.M., Gotter, A.L., Roecker, A.J., Winrow, C.J., Renger, J.J., Coleman, P.J., Rosenbaum, D.M.(2016) Nat Struct Mol Biol 23: 293-299
- PubMed: 26950369 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.3183
- Primary Citation Related Structures:
4ZJ8, 4ZJC - PubMed Abstract:
The orexin (also known as hypocretin) G protein-coupled receptors (GPCRs) regulate sleep and other behavioral functions in mammals, and are therapeutic targets for sleep and wake disorders. The human receptors hOX1R and hOX2R, which are 64% identical in sequence, have overlapping but distinct physiological functions and potential therapeutic profiles. We determined structures of hOX1R bound to the OX1R-selective antagonist SB-674042 and the dual antagonist suvorexant at 2.8-Å and 2.75-Å resolution, respectively, and used molecular modeling to illuminate mechanisms of antagonist subtype selectivity between hOX1R and hOX2R. The hOX1R structures also reveal a conserved amphipathic α-helix, in the extracellular N-terminal region, that interacts with orexin-A and is essential for high-potency neuropeptide activation at both receptors. The orexin-receptor crystal structures are valuable tools for the design and development of selective orexin-receptor antagonists and agonists.
- Department of Biophysics, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
Organizational Affiliation: