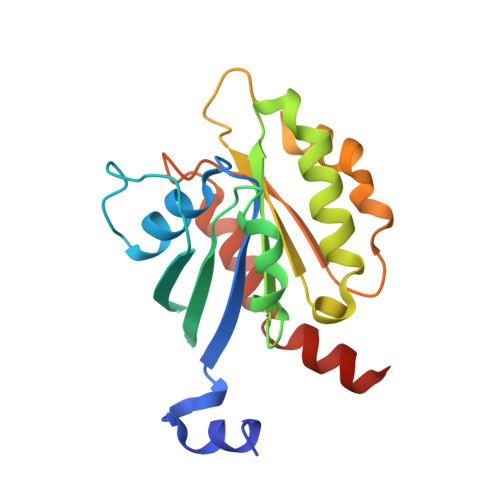
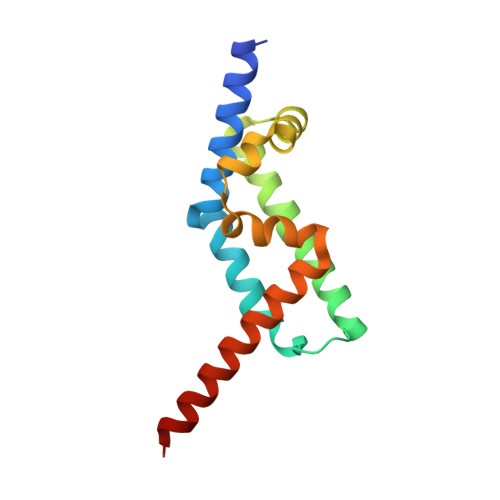
The Interaction of CCDC104/BARTL1 with Arl3 and Implications for Ciliary Function.
Lokaj, M., Kosling, S.K., Koerner, C., Lange, S.M., van Beersum, S.E., van Reeuwijk, J., Roepman, R., Horn, N., Ueffing, M., Boldt, K., Wittinghofer, A.(2015) Structure 23: 2122-2132
- PubMed: 26455799 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2015.08.016
- Primary Citation Related Structures:
4ZI2, 4ZI3 - PubMed Abstract:
Cilia are small antenna-like cellular protrusions critical for many developmental signaling pathways. The ciliary protein Arl3 has been shown to act as a specific release factor for myristoylated and farnesylated ciliary cargo molecules by binding to the effectors Unc119 and PDE6δ. Here we describe a newly identified Arl3 binding partner, CCDC104/CFAP36. Biochemical and structural analyses reveal that the protein contains a BART-like domain and is called BARTL1. It recognizes an LLxILxxL motif at the N-terminal amphipathic helix of Arl3, which is crucial for the interaction with the BART-like domain but also for the ciliary localization of Arl3 itself. These results seem to suggest a ciliary role of BARTL1, and possibly link it to the Arl3 transport network. We thus speculate on a regulatory mechanism whereby BARTL1 aids the presentation of active Arl3 to its GTPase-activating protein RP2 or hinders Arl3 membrane binding in the area of the transition zone.
- Max-Planck-Institute of Molecular Physiology, Emeritus Group, Otto-Hahn-Straße 15, 44227 Dortmund, Germany.
Organizational Affiliation: