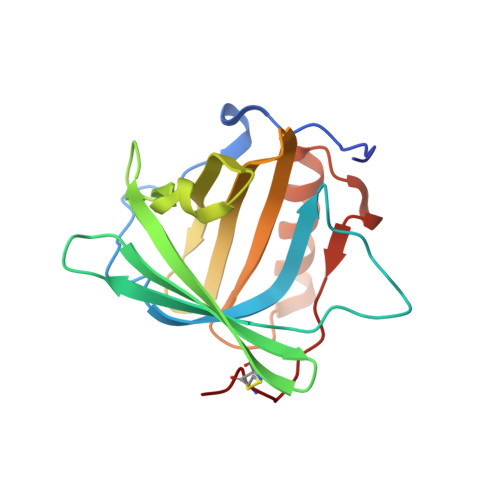
Siderocalin-mediated recognition, sensitization, and cellular uptake of actinides.
Allred, B.E., Rupert, P.B., Gauny, S.S., An, D.D., Ralston, C.Y., Sturzbecher-Hoehne, M., Strong, R.K., Abergel, R.J.(2015) Proc Natl Acad Sci U S A 112: 10342-10347
- PubMed: 26240330 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1508902112
- Primary Citation Related Structures:
4ZFX, 4ZHC, 4ZHD, 4ZHF, 4ZHG, 4ZHH - PubMed Abstract:
Synthetic radionuclides, such as the transuranic actinides plutonium, americium, and curium, present severe health threats as contaminants, and understanding the scope of the biochemical interactions involved in actinide transport is instrumental in managing human contamination. Here we show that siderocalin, a mammalian siderophore-binding protein from the lipocalin family, specifically binds lanthanide and actinide complexes through molecular recognition of the ligands chelating the metal ions. Using crystallography, we structurally characterized the resulting siderocalin-transuranic actinide complexes, providing unprecedented insights into the biological coordination of heavy radioelements. In controlled in vitro assays, we found that intracellular plutonium uptake can occur through siderocalin-mediated endocytosis. We also demonstrated that siderocalin can act as a synergistic antenna to sensitize the luminescence of trivalent lanthanide and actinide ions in ternary protein-ligand complexes, dramatically increasing the brightness and efficiency of intramolecular energy transfer processes that give rise to metal luminescence. Our results identify siderocalin as a potential player in the biological trafficking of f elements, but through a secondary ligand-based metal sequestration mechanism. Beyond elucidating contamination pathways, this work is a starting point for the design of two-stage biomimetic platforms for photoluminescence, separation, and transport applications.
- Chemical Sciences Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720;
Organizational Affiliation: