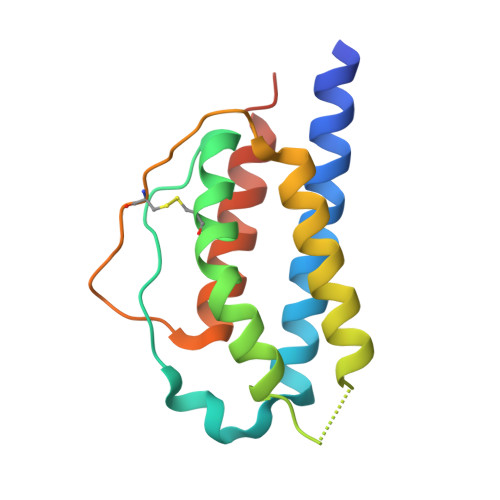
Structural and functional characterisation of ferret interleukin-2.
Ren, B., McKinstry, W.J., Pham, T., Newman, J., Layton, D.S., Bean, A.G., Chen, Z., Laurie, K.L., Borg, K., Barr, I.G., Adams, T.E.(2015) Dev Comp Immunol 55: 32-38
- PubMed: 26472619 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.dci.2015.10.007
- Primary Citation Related Structures:
4ZF7 - PubMed Abstract:
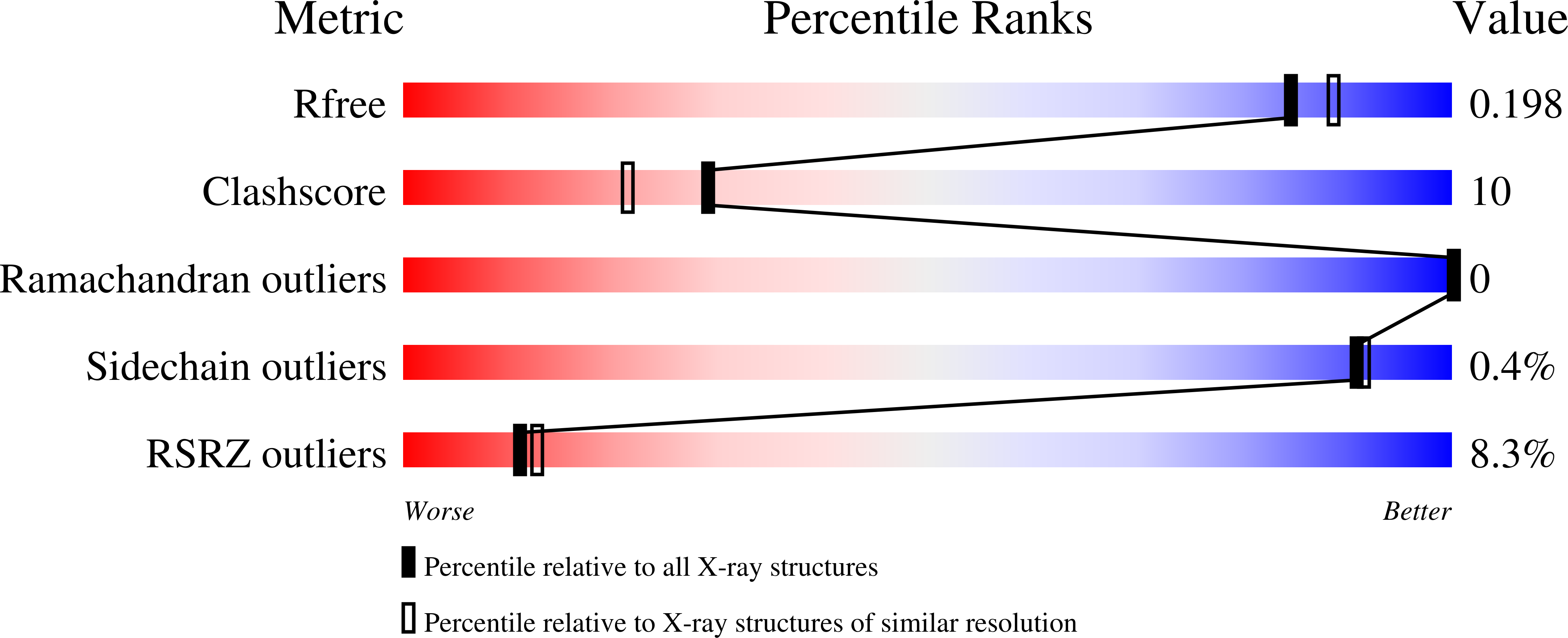
While the ferret is a valuable animal model for a number of human viral infections, such as influenza, Hendra and Nipah, evaluating the cellular immune response following infection has been hampered by the lack of a number of species-specific immunological reagents. Interleukin 2 (IL-2) is one such key cytokine. Ferret recombinant IL-2 incorporating a C-terminal histidine tag was expressed and purified and the three-dimensional structure solved and refined at 1.89 Å by X-ray crystallography, which represents the highest resolution and first non-human IL-2 structure. While ferret IL-2 displays the classic cytokine fold of the four-helix bundle structure, conformational flexibility was observed at the second helix and its neighbouring region in the bundle, which may result in the disruption of the spatial arrangement of residues involved in receptor binding interactions, implicating subtle differences between ferret and human IL-2 when initiating biological functions. Ferret recombinant IL-2 stimulated the proliferation of ferret lymph node cells and induced the expression of mRNA for IFN-γ and Granzyme A.
- CSIRO Manufacturing, Parkville, VIC 3052, Australia.
Organizational Affiliation: