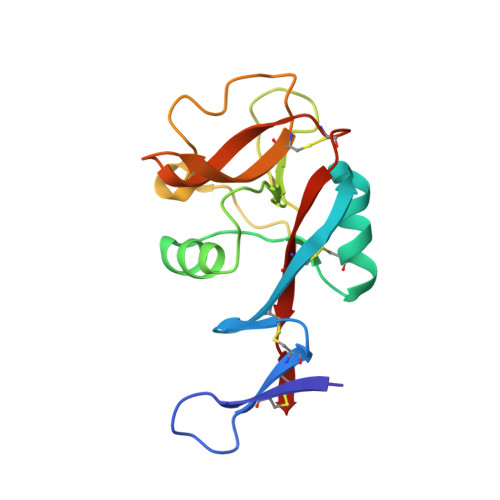
A Novel Mechanism for Binding of Galactose-terminated Glycans by the C-type Carbohydrate Recognition Domain in Blood Dendritic Cell Antigen 2.
Jegouzo, S.A., Feinberg, H., Dungarwalla, T., Drickamer, K., Weis, W.I., Taylor, M.E.(2015) J Biological Chem 290: 16759-16771
- PubMed: 25995448 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.660613
- Primary Citation Related Structures:
4ZES, 4ZET - PubMed Abstract:
Blood dendritic cell antigen 2 (BDCA-2; also designated CLEC4C or CD303) is uniquely expressed on plasmacytoid dendritic cells. Stimulation of BDCA-2 with antibodies leads to an anti-inflammatory response in these cells, but the natural ligands for the receptor are not known. The C-type carbohydrate recognition domain in the extracellular portion of BDCA-2 contains a signature motif typical of C-type animal lectins that bind mannose, glucose, or GlcNAc, yet it has been reported that BDCA-2 binds selectively to galactose-terminated, biantennary N-linked glycans. A combination of glycan array analysis and binding competition studies with monosaccharides and natural and synthetic oligosaccharides have been used to define the binding epitope for BDCA-2 as the trisaccharide Galβ1-3/4GlcNAcβ1-2Man. X-ray crystallography and mutagenesis studies show that mannose is ligated to the conserved Ca(2+) in the primary binding site that is characteristic of C-type carbohydrate recognition domains, and the GlcNAc and galactose residues make additional interactions in a wide, shallow groove adjacent to the primary binding site. As predicted from these studies, BDCA-2 binds to IgG, which bears galactose-terminated glycans that are not commonly found attached to other serum glycoproteins. Thus, BDCA-2 has the potential to serve as a previously unrecognized immunoglobulin Fc receptor.
- the Department of Life Sciences, Imperial College, London SW7 2AZ, United Kingdom.
Organizational Affiliation: