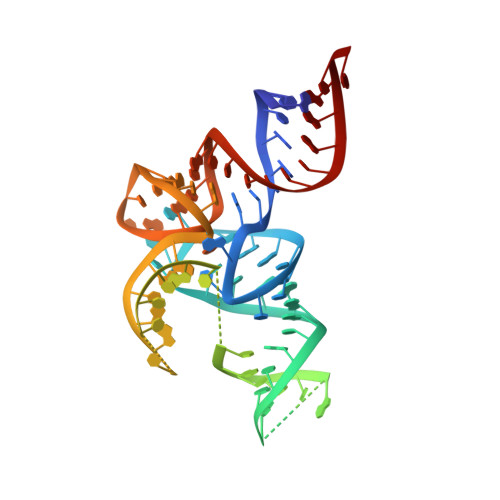
Structural basis for early-onset neurological disorders caused by mutations in human selenocysteine synthase.
Puppala, A.K., French, R.L., Matthies, D., Baxa, U., Subramaniam, S., Simonovic, M.(2016) Sci Rep 6: 32563-32563
- PubMed: 27576344 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep32563
- Primary Citation Related Structures:
4ZDL, 4ZDO, 4ZDP - PubMed Abstract:
Selenocysteine synthase (SepSecS) catalyzes the terminal reaction of selenocysteine, and is vital for human selenoproteome integrity. Autosomal recessive inheritance of mutations in SepSecS-Ala239Thr, Thr325Ser, Tyr334Cys and Tyr429*-induced severe, early-onset, neurological disorders in distinct human populations. Although harboring different mutant alleles, patients presented remarkably similar phenotypes typified by cerebellar and cerebral atrophy, seizures, irritability, ataxia, and extreme spasticity. However, it has remained unclear how these genetic alterations affected the structure of SepSecS and subsequently elicited the development of a neurological pathology. Herein, our biophysical and structural characterization demonstrates that, with the exception of Tyr429*, pathogenic mutations decrease protein stability and trigger protein misfolding. We propose that the reduced stability and increased propensity towards misfolding are the main causes for the loss of SepSecS activity in afflicted patients, and that these factors contribute to disease progression. We also suggest that misfolding of enzymes regulating protein synthesis should be considered in the diagnosis and study of childhood neurological disorders.
- Department of Biochemistry an Molecular Genetics, University of Illinois at Chicago, Chicago, Illinois 60607, USA.
Organizational Affiliation: