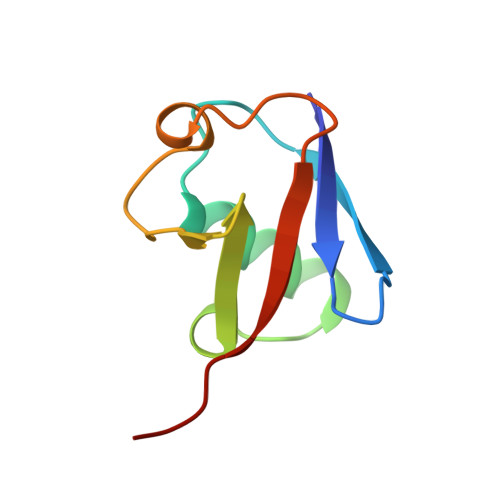
Tetrameric Assembly of Monoubiquitin Accurately Mimics the Lys11 Polyubiquitin Chain Structure.
Levin-Kravets, O., Shohat, N., Prag, G.(2015) Biochemistry 54: 4704-4710
- PubMed: 26171660 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.5b00498
- Primary Citation Related Structures:
4Z9S - PubMed Abstract:
Specific lysine residues on the ubiquitin surface were selected during the course of evolution to form different polyubiquitin chain structures that signal diverse cellular processes. A vast number of ubiquitin receptors specifically recognize and decode the signals conferred by these polyubiquitin chains. The mechanisms of formation and the structure of Lys11-linked ubiquitin, which signals for cell-cycle and innate immune control, have been elucidated. Here, we present a new crystal structure of monomeric ubiquitin that accurately mimics one of the structures of Lys11-linked ubiquitin. Analysis of the ubiquitin:ubiquitin interface demonstrates structural fitness and specificity. The interaction is exclusively hydrophilic, leaving the Ile44 hydrophobic patch, a major recognition site for ubiquitin receptors, exposed. These noncovalent ubiquitin:ubiquitin interactions are nearly identical to those reported for Lys11-linked ubiquitin and seem to play a significant role in stabilizing the crystal structure without the isopeptide bond. In vitro cross-linking analysis with wild-type ubiquitin or its mutants partially mimics the interactions in the crystal. We suggest that these interactions may play a biological role in transmitting Lys11-linked ubiquitin chain-type cellular signals.
- Department of Biochemistry and Molecular Biology and Institute for Structural Biology, George S. Wise Faculty of Life Sciences, Tel Aviv University, Tel Aviv, Israel.
Organizational Affiliation: