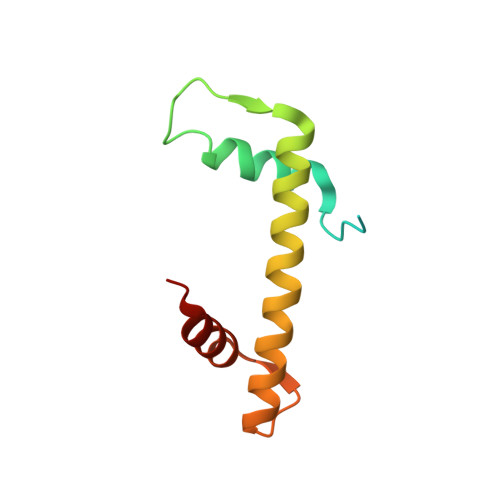
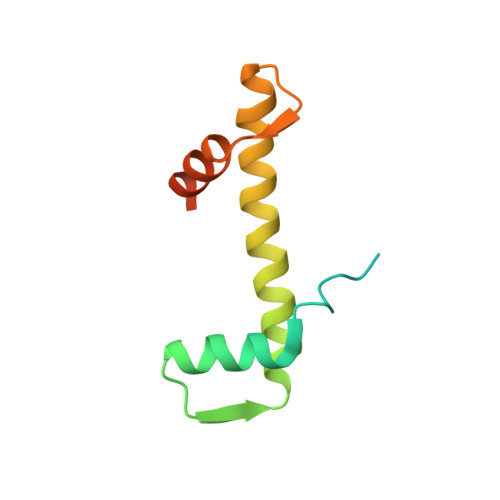
Integrated molecular mechanism directing nucleosome reorganization by human FACT.
Tsunaka, Y., Fujiwara, Y., Oyama, T., Hirose, S., Morikawa, K.(2016) Genes Dev 30: 673-686
- PubMed: 26966247 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/gad.274183.115
- Primary Citation Related Structures:
4Z2M, 4Z2N - PubMed Abstract:
Facilitates chromatin transcription (FACT) plays essential roles in chromatin remodeling during DNA transcription, replication, and repair. Our structural and biochemical studies of human FACT-histone interactions present precise views of nucleosome reorganization, conducted by the FACT-SPT16 (suppressor of Ty 16) Mid domain and its adjacent acidic AID segment. AID accesses the H2B N-terminal basic region exposed by partial unwrapping of the nucleosomal DNA, thereby triggering the invasion of FACT into the nucleosome. The crystal structure of the Mid domain complexed with an H3-H4 tetramer exhibits two separate contact sites; the Mid domain forms a novel intermolecular β structure with H4. At the other site, the Mid-H2A steric collision on the H2A-docking surface of the H3-H4 tetramer within the nucleosome induces H2A-H2B displacement. This integrated mechanism results in disrupting the H3 αN helix, which is essential for retaining the nucleosomal DNA ends, and hence facilitates DNA stripping from histone.
- Precursory Research for Embryonic Science and Technology (PRESTO), Japan Science and Technology Agency, Sakyo-ku, Kyoto 606-8501, Japan; Institute for Integrated Cell-Material Sciences (iCeMS), Kyoto University, Sakyo-ku, Kyoto 606-8501, Japan; Department of Gene Mechanisms, Graduate School of Biostudies, Kyoto University, Yoshida-konoemachi, Sakyo-ku, Kyoto 606-8501, Japan; International Institute for Advanced Studies, Kizugawa-shi, Kyoto 619-0225, Japan;
Organizational Affiliation: