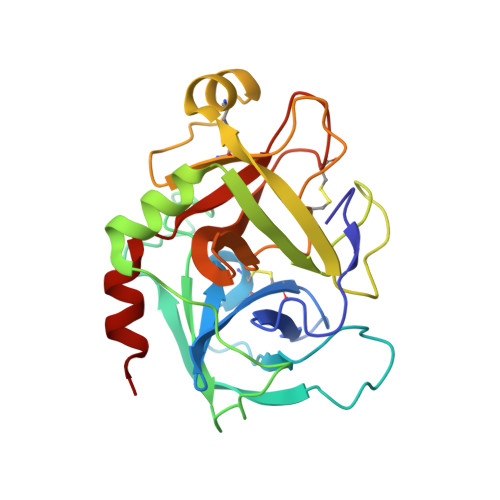
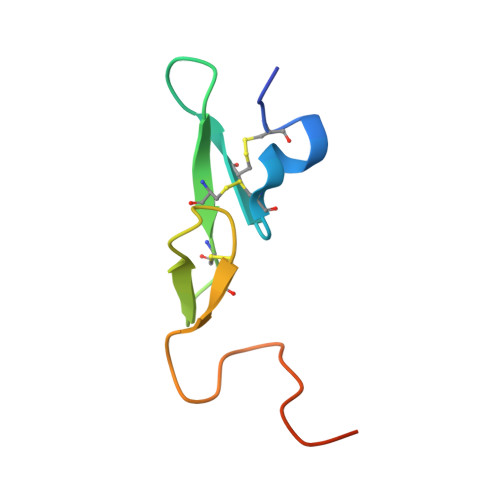
Rapid development of two factor IXa inhibitors from hit to lead.
Parker, D.L., Walsh, S., Li, B., Kim, E., Sharipour, A., Smith, C., Chen, Y.H., Berger, R., Harper, B., Zhang, T., Park, M., Shu, M., Wu, J., Xu, J., Dewnani, S., Sherer, E.C., Hruza, A., Reichert, P., Geissler, W., Sonatore, L., Ellsworth, K., Balkovec, J., Greenlee, W., Wood, H.B.(2015) Bioorg Med Chem Lett 25: 2321-2325
- PubMed: 25937013 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2015.04.025
- Primary Citation Related Structures:
4YZU, 4Z0K - PubMed Abstract:
Two high-throughput screening hits were investigated for SAR against human factor IXa. Both hits feature a benzamide linked to a [6-5]-heteroaryl via an alkyl amine. In the case where this system is a benzimidazolyl-ethyl amine the binding potency for the hit was improved >500-fold, from 9 μM to 0.016 μM. For the other hit, which contains a tetrahydropyrido-indazole amine, potency was improved 20-fold, from 2 μM to 0.09 μM. X-ray crystal structures were obtained for an example of each class which improved understanding of the binding, and will enable further drug discovery efforts.
- Department of Medicinal Chemistry, Merck Research Laboratories, Merck and Co., Inc, PO Box 2000, E. Lincoln Ave., Rahway, NJ 07016, USA. Electronic address: dann_parker@merck.com.
Organizational Affiliation: