Structure of hemagglutinin from a H6N1 influenza virus (A/Taiwan/2/2013) at 2.6 Angstroms resolution
Wang, F., Qi, J., Bi, Y., Zhang, W., Wang, M., Wang, M., Liu, J., Yan, J., Shi, Y., Gao, G.F.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| HA1 | 325 | H6N1 subtype | Mutation(s): 0 | 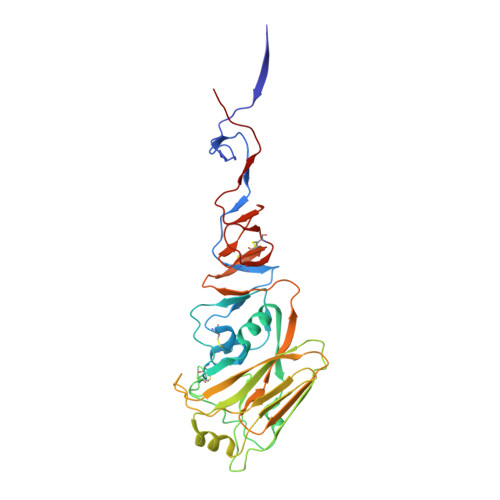 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A0J9X268 | ||||
Glycosylation | |||||
| Glycosylation Sites: 2 | |||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| HA2 | 162 | H6N1 subtype | Mutation(s): 0 |  | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A0J9X267 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | |||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| N-acetyl-alpha-neuraminic acid-(2-6)-beta-D-galactopyranose | C | 2 |  | N/A | |
Glycosylation Resources | |||||
GlyTouCan: G63069TR GlyCosmos: G63069TR GlyGen: G63069TR | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Download:Ideal Coordinates CCD File | E [auth A], F [auth A] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 114.387 | α = 90 |
| b = 114.387 | β = 90 |
| c = 166.049 | γ = 120 |
| Software Name | Purpose |
|---|---|
| HKL-2000 | data processing |
| PHASER | phasing |
| Coot | model building |
| PHENIX | refinement |
| SCALA | data reduction |
| SCALEPACK | data scaling |
| Funding Organization | Location | Grant Number |
|---|---|---|
| China Ministry of Science and Technology National 973 Project | China | 2011CB504703 |
| Intramural Special Grant for Influenza Virus Research from the Chinese Academy of Sciences | China | KJZD-EW-L09 |
| Intramural Special Grant for Strategic Priority Research Program of the Chinese Academy of Sciences | China | XDB08020100 |
| National Natural Science Foundation of China | China | 31402196 |
| China National Grand S&T Special Project | China | 2014ZX10004002 |