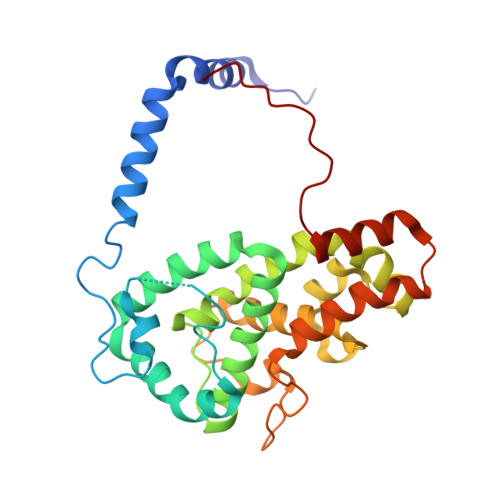
The structure of the folded domain from the signature multifunctional protein ICP27 from herpes simplex virus-1 reveals an intertwined dimer.
Tunnicliffe, R.B., Schacht, M., Levy, C., Jowitt, T.A., Sandri-Goldin, R.M., Golovanov, A.P.(2015) Sci Rep 5: 11234-11234
- PubMed: 26062451 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep11234
- Primary Citation Related Structures:
4YXP - PubMed Abstract:
Herpesviruses cause life-long infections by evading the host immune system and establishing latent infections. All mammalian herpesviruses express an essential multifunctional protein that is typified by ICP27 encoded by Herpes Simplex Virus 1. The only region that is conserved among the diverse members of the ICP27 family is a predicted globular domain that has been termed the ICP27 homology domain. Here we present the first crystal structure of the ICP27 homology domain, solved to 1.9 Å resolution. The protein is a homo-dimer, adopting a novel intertwined fold with one CHCC zinc-binding site per monomer. The dimerization, which was independently confirmed by SEC-MALS and AUC, is stabilized by an extensive network of intermolecular contacts, and a domain-swap involving the two N-terminal helices and C-terminal tails. Each monomer contains a lid motif that can clamp the C-terminal tail of its dimeric binding partner against its globular core, without forming any distinct secondary structure elements. The binding interface was probed with point mutations, none of which had a noticeable effect on dimer formation; however deletion of the C-terminal tail region prevented dimer formation in vivo. The structure provides a template for future biochemical studies and modelling of ICP27 homologs from other herpesviruses.
- Manchester Institute of Biotechnology, The University of Manchester, Manchester, UK.
Organizational Affiliation: