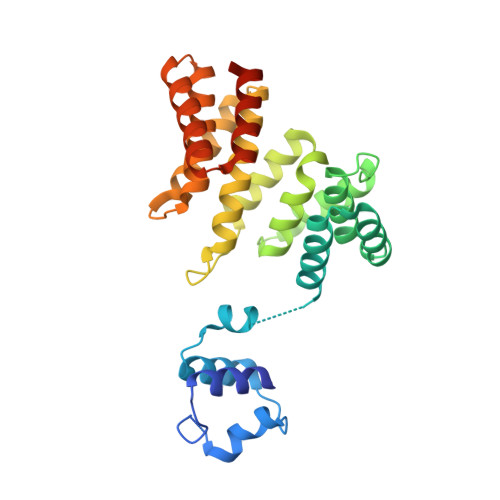
Rgg protein structure-function and inhibition by cyclic peptide compounds.
Parashar, V., Aggarwal, C., Federle, M.J., Neiditch, M.B.(2015) Proc Natl Acad Sci U S A 112: 5177-5182
- PubMed: 25847993 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1500357112
- Primary Citation Related Structures:
4YV6, 4YV9 - PubMed Abstract:
Peptide pheromone cell-cell signaling (quorum sensing) regulates the expression of diverse developmental phenotypes (including virulence) in Firmicutes, which includes common human pathogens, e.g., Streptococcus pyogenes and Streptococcus pneumoniae. Cytoplasmic transcription factors known as "Rgg proteins" are peptide pheromone receptors ubiquitous in Firmicutes. Here we present X-ray crystal structures of a Streptococcus Rgg protein alone and in complex with a tight-binding signaling antagonist, the cyclic undecapeptide cyclosporin A. To our knowledge, these represent the first Rgg protein X-ray crystal structures. Based on the results of extensive structure-function analysis, we reveal the peptide pheromone-binding site and the mechanism by which cyclosporin A inhibits activation of the peptide pheromone receptor. Guided by the Rgg-cyclosporin A complex structure, we predicted that the nonimmunosuppressive cyclosporin A analog valspodar would inhibit Rgg activation. Indeed, we found that, like cyclosporin A, valspodar inhibits peptide pheromone activation of conserved Rgg proteins in medically relevant Streptococcus species. Finally, the crystal structures presented here revealed that the Rgg protein DNA-binding domains are covalently linked across their dimerization interface by a disulfide bond formed by a highly conserved cysteine. The DNA-binding domain dimerization interface observed in our structures is essentially identical to the interfaces previously described for other members of the XRE DNA-binding domain family, but the presence of an intermolecular disulfide bond buried in this interface appears to be unique. We hypothesize that this disulfide bond may, under the right conditions, affect Rgg monomer-dimer equilibrium, stabilize Rgg conformation, or serve as a redox-sensitive switch.
- Department of Microbiology, Biochemistry, and Molecular Genetics, New Jersey Medical School, Rutgers, The State University of New Jersey, Newark, NJ 07103; and.
Organizational Affiliation: