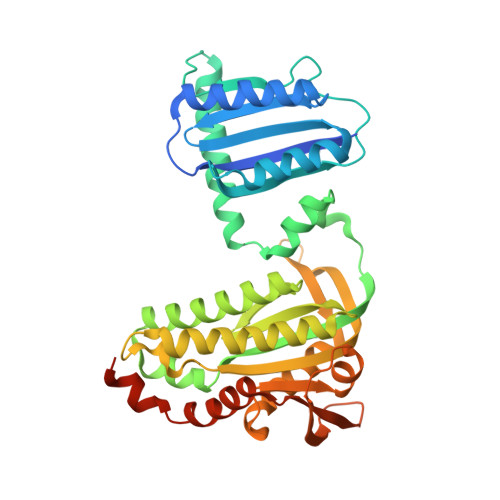
Structural insight into photoactivation of an adenylate cyclase from a photosynthetic cyanobacterium
Ohki, M., Sugiyama, K., Kawai, F., Tanaka, H., Nihei, Y., Unzai, S., Takebe, M., Matsunaga, S., Adachi, S., Shibayama, N., Zhou, Z., Koyama, R., Ikegaya, Y., Takahashi, T., Tame, J.R.H., Iseki, M., Park, S.-Y.(2016) Proc Natl Acad Sci U S A 113: 6659-6664
- PubMed: 27247413 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1517520113
- Primary Citation Related Structures:
4YUS, 4YUT - PubMed Abstract:
Cyclic-AMP is one of the most important second messengers, regulating many crucial cellular events in both prokaryotes and eukaryotes, and precise spatial and temporal control of cAMP levels by light shows great promise as a simple means of manipulating and studying numerous cell pathways and processes. The photoactivated adenylate cyclase (PAC) from the photosynthetic cyanobacterium Oscillatoria acuminata (OaPAC) is a small homodimer eminently suitable for this task, requiring only a simple flavin chromophore within a blue light using flavin (BLUF) domain. These domains, one of the most studied types of biological photoreceptor, respond to blue light and either regulate the activity of an attached enzyme domain or change its affinity for a repressor protein. BLUF domains were discovered through studies of photo-induced movements of Euglena gracilis, a unicellular flagellate, and gene expression in the purple bacterium Rhodobacter sphaeroides, but the precise details of light activation remain unknown. Here, we describe crystal structures and the light regulation mechanism of the previously undescribed OaPAC, showing a central coiled coil transmits changes from the light-sensing domains to the active sites with minimal structural rearrangement. Site-directed mutants show residues essential for signal transduction over 45 Å across the protein. The use of the protein in living human cells is demonstrated with cAMP-dependent luciferase, showing a rapid and stable response to light over many hours and activation cycles. The structures determined in this study will assist future efforts to create artificial light-regulated control modules as part of a general optogenetic toolkit.
- Drug Design Laboratory, Graduate School of Medical Life Science, Yokohama City University, Tsurumi, Yokohama, 230-0045, Japan;
Organizational Affiliation: