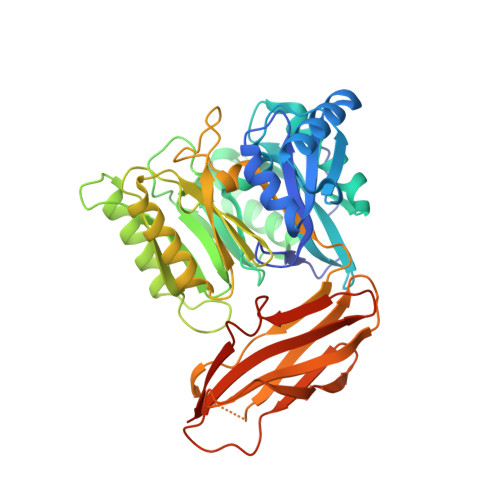
Structure and mechanism of a bacterial host-protein citrullinating virulence factor, Porphyromonas gingivalis peptidylarginine deiminase.
Goulas, T., Mizgalska, D., Garcia-Ferrer, I., Kantyka, T., Guevara, T., Szmigielski, B., Sroka, A., Millan, C., Uson, I., Veillard, F., Potempa, B., Mydel, P., Sola, M., Potempa, J., Gomis-Ruth, F.X.(2015) Sci Rep 5: 11969-11969
- PubMed: 26132828 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep11969
- Primary Citation Related Structures:
4YT9, 4YTB, 4YTG - PubMed Abstract:
Citrullination is a post-translational modification of higher organisms that deiminates arginines in proteins and peptides. It occurs in physiological processes but also pathologies such as multiple sclerosis, fibrosis, Alzheimer's disease and rheumatoid arthritis (RA). The reaction is catalyzed by peptidylarginine deiminases (PADs), which are found in vertebrates but not in lower organisms. RA has been epidemiologically associated with periodontal disease, whose main infective agent is Porphyromonas gingivalis. Uniquely among microbes, P. gingivalis secretes a PAD, termed PPAD (Porphyromonas peptidylarginine deiminase), which is genetically unrelated to eukaryotic PADs. Here, we studied function of PPAD and its substrate-free, substrate-complex, and substrate-mimic-complex structures. It comprises a flat cylindrical catalytic domain with five-fold α/β-propeller architecture and a C-terminal immunoglobulin-like domain. The PPAD active site is a funnel located on one of the cylinder bases. It accommodates arginines from peptide substrates after major rearrangement of a "Michaelis loop" that closes the cleft. The guanidinium and carboxylate groups of substrates are tightly bound, which explains activity of PPAD against arginines at C-termini but not within peptides. Catalysis is based on a cysteine-histidine-asparagine triad, which is shared with human PAD1-PAD4 and other guanidino-group modifying enzymes. We provide a working mechanism hypothesis based on 18 structure-derived point mutants.
- Proteolysis Lab; Department of Structural Biology ("María de Maeztu" Unit of Excellence); Molecular Biology Institute of Barcelona, CSIC; Barcelona Science Park, Helix Building; c/Baldiri Reixac, 15-21; E-08028 Barcelona Spain.
Organizational Affiliation: