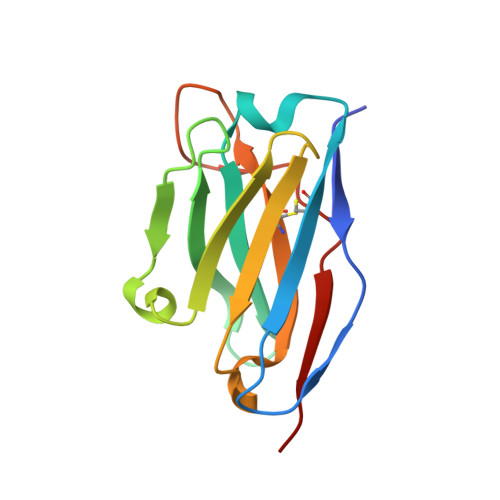
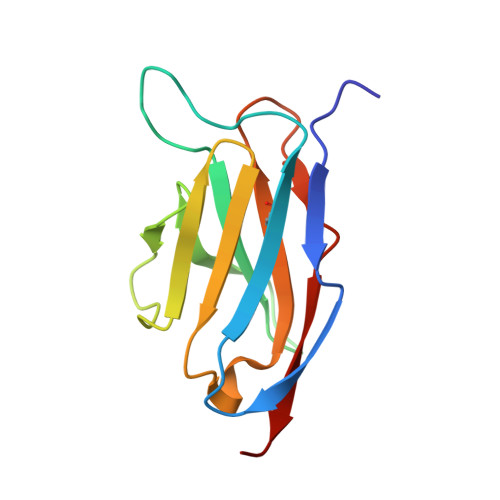
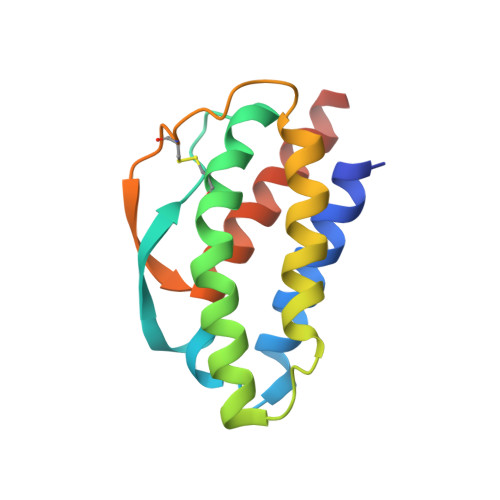
Antibodies to Interleukin-2 Elicit Selective T Cell Subset Potentiation through Distinct Conformational Mechanisms.
Spangler, J.B., Tomala, J., Luca, V.C., Jude, K.M., Dong, S., Ring, A.M., Votavova, P., Pepper, M., Kovar, M., Garcia, K.C.(2015) Immunity 42: 815-825
- PubMed: 25992858 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.immuni.2015.04.015
- Primary Citation Related Structures:
4YQX, 4YUE - PubMed Abstract:
Interleukin-2 (IL-2) is a pleiotropic cytokine that regulates immune cell homeostasis and has been used to treat a range of disorders including cancer and autoimmune disease. IL-2 signals via interleukin-2 receptor-β (IL-2Rβ):IL-2Rγ heterodimers on cells expressing high (regulatory T cells, Treg) or low (effector cells) amounts of IL-2Rα (CD25). When complexed with IL-2, certain anti-cytokine antibodies preferentially stimulate expansion of Treg (JES6-1) or effector (S4B6) cells, offering a strategy for targeted disease therapy. We found that JES6-1 sterically blocked the IL-2:IL-2Rβ and IL-2:IL-2Rγ interactions, but also allosterically lowered the IL-2:IL-2Rα affinity through a "triggered exchange" mechanism favoring IL-2Rα(hi) Treg cells, creating a positive feedback loop for IL-2Rα(hi) cell activation. Conversely, S4B6 sterically blocked the IL-2:IL-2Rα interaction, while also conformationally stabilizing the IL-2:IL-2Rβ interaction, thus stimulating all IL-2-responsive immune cells, particularly IL-2Rβ(hi) effector cells. These insights provide a molecular blueprint for engineering selectively potentiating therapeutic antibodies.
- Howard Hughes Medical Institute, Stanford University School of Medicine, Stanford, CA 94305, USA; Department of Molecular and Cellular Physiology, Stanford University School of Medicine, Stanford, CA 94305, USA; Department of Structural Biology, Stanford University School of Medicine, Stanford, CA 94305, USA.
Organizational Affiliation: