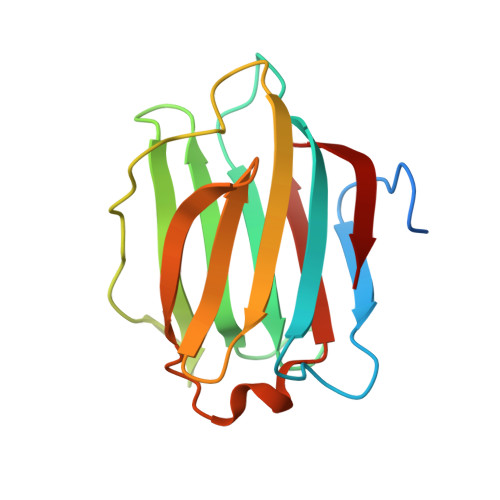
Structural characterization of human galectin-4 C-terminal domain: elucidating the molecular basis for recognition of glycosphingolipids, sulfated saccharides and blood group antigens.
Bum-Erdene, K., Leffler, H., Nilsson, U.J., Blanchard, H.(2015) FEBS J 282: 3348-3367
- PubMed: 26077389 Search on PubMed
- DOI: https://doi.org/10.1111/febs.13348
- Primary Citation Related Structures:
4YLZ, 4YM0, 4YM1, 4YM2, 4YM3 - PubMed Abstract:
Human galectin-4 is a lectin that is expressed mainly in the gastrointestinal tract and exhibits metastasis-promoting roles in some cancers. Its tandem-repeat nature exhibits two distinct carbohydrate recognition domains allowing crosslinking by simultaneous binding to sulfated and non-sulfated (but not sialylated) glycosphingolipids and glycoproteins, facilitating stabilization of lipid rafts. Critically, galectin-4 exerts favourable or unfavourable effects depending upon the cancer. Here we report the first X-ray crystallographic structural information on human galectin-4, specifically the C-terminal carbohydrate recognition domain of human (galectin-4C) in complex with lactose, lactose-3'-sulfate, 2'-fucosyllactose, lacto-N-tetraose and lacto-N-neotetraose. These structures enable elucidation of galectin-4C binding fine-specificity towards sulfated and non-sulfated lacto- and neolacto-series sphingolipids as well as to human blood group antigens. Analysis of the lactose-3'-sulfate complex structure shows that galectin-4C does not recognize the sulfate group using any specific amino acid, but binds the ligand nonetheless. Complex structures with lacto-N-tetraose and lacto-N-neotetraose displayed differences in binding interactions exhibited by the non-reducing-end galactose. That of lacto-N-tetraose points outward from the protein surface whereas that of lacto-N-neotetraose interacts directly with the protein. Recognition patterns of human galectin-4C towards lacto- and neolacto-series glycosphingolipids are similar to those of human galectin-3; however, detailed scrutiny revealed differences stemming from the extended binding site that offer distinction in ligand profiles of these two galectins. Structural characterization of the complex with 2'-fucosyllactose, a carbohydrate with similarity to the H antigen, and molecular dynamics studies highlight structural features that allow specific recognition of A and B antigens, whilst a lack of interaction with the 2'-fucose of blood group antigens was revealed. 4YLZ, 4YM0, 4YM1, 4YM2, 4YM3.
- Institute for Glycomics, Griffith University, Australia.
Organizational Affiliation: