Discovery of a Potent Class of PI3K alpha Inhibitors with Unique Binding Mode via Encoded Library Technology (ELT).
Yang, H., Medeiros, P.F., Raha, K., Elkins, P., Lind, K.E., Lehr, R., Adams, N.D., Burgess, J.L., Schmidt, S.J., Knight, S.D., Auger, K.R., Schaber, M.D., Franklin, G.J., Ding, Y., DeLorey, J.L., Centrella, P.A., Mataruse, S., Skinner, S.R., Clark, M.A., Cuozzo, J.W., Evindar, G.(2015) ACS Med Chem Lett 6: 531-536
- PubMed: 26005528 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.5b00025
- Primary Citation Related Structures:
4YKN - PubMed Abstract:
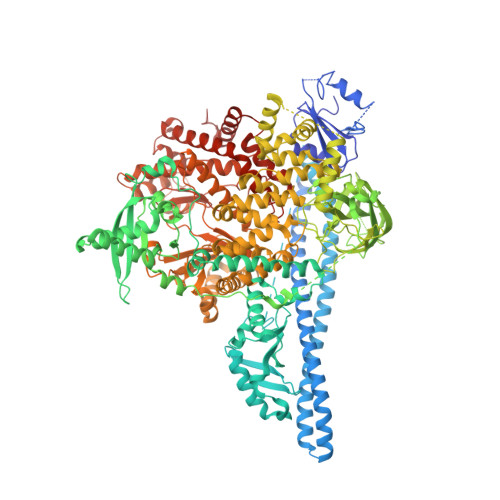
In the search of PI3K p110α wild type and H1047R mutant selective small molecule leads, an encoded library technology (ELT) campaign against the desired target proteins was performed which led to the discovery of a selective chemotype for PI3K isoforms from a three-cycle DNA encoded library. An X-ray crystal structure of a representative inhibitor from this chemotype demonstrated a unique binding mode in the p110α protein.
- MDR (Molecular Discovery Research) Boston, Platform Technology and Science, GlaxoSmithKline , 830 Winter Street, Waltham, Massachusetts 02451, United States.
Organizational Affiliation: