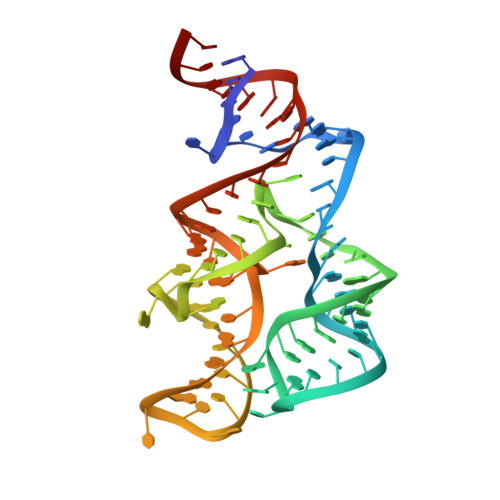
Structural Basis for Molecular Discrimination by a 3',3'-cGAMP Sensing Riboswitch.
Ren, A., Wang, X.C., Kellenberger, C.A., Rajashankar, K.R., Jones, R.A., Hammond, M.C., Patel, D.J.(2015) Cell Rep 11: 1-12
- PubMed: 25818298 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2015.03.004
- Primary Citation Related Structures:
4YAZ, 4YB0, 4YB1 - PubMed Abstract:
Cyclic dinucleotides are second messengers that target the adaptor STING and stimulate the innate immune response in mammals. Besides protein receptors, there are bacterial riboswitches that selectively recognize cyclic dinucleotides. We recently discovered a natural riboswitch that targets 3',3'-cGAMP, which is distinguished from the endogenous mammalian signal 2',3'-cGAMP by its backbone connectivity. Here, we report on structures of the aptamer domain of the 3',3'-cGAMP riboswitch from Geobacter in the 3',3'-cGAMP and c-di-GMP bound states. The riboswitch adopts a tuning fork-like architecture with a junctional ligand-binding pocket and different orientations of the arms are correlated with the identity of the bound cyclic dinucleotide. Subsequent biochemical experiments revealed that specificity of ligand recognition can be affected by point mutations outside of the binding pocket, which has implications for both the assignment and reengineering of riboswitches in this structural class.
- Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, NY 10065, USA.
Organizational Affiliation: