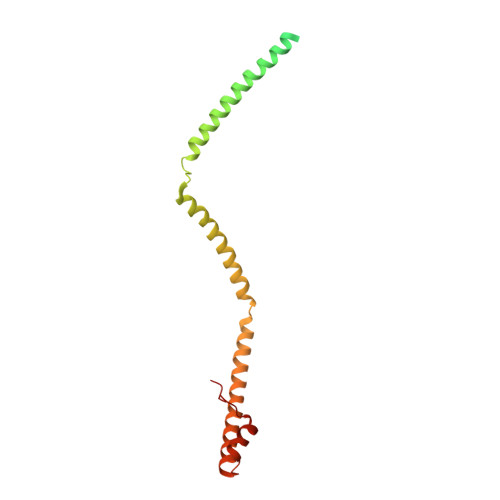
Crystal structure of Hop2-Mnd1 and mechanistic insights into its role in meiotic recombination
Kang, H.A., Shin, H.C., Kalantzi, A.S., Toseland, C.P., Kim, H.M., Gruber, S., Peraro, M.D., Oh, B.H.(2015) Nucleic Acids Res 43: 3841-3856
- PubMed: 25740648 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkv172
- Primary Citation Related Structures:
4Y66 - PubMed Abstract:
In meiotic DNA recombination, the Hop2-Mnd1 complex promotes Dmc1-mediated single-stranded DNA (ssDNA) invasion into homologous chromosomes to form a synaptic complex by a yet-unclear mechanism. Here, the crystal structure of Hop2-Mnd1 reveals that it forms a curved rod-like structure consisting of three leucine zippers and two kinked junctions. One end of the rod is linked to two juxtaposed winged-helix domains, and the other end is capped by extra α-helices to form a helical bundle-like structure. Deletion analysis shows that the helical bundle-like structure is sufficient for interacting with the Dmc1-ssDNA nucleofilament, and molecular modeling suggests that the curved rod could be accommodated into the helical groove of the nucleofilament. Remarkably, the winged-helix domains are juxtaposed at fixed relative orientation, and their binding to DNA is likely to perturb the base pairing according to molecular simulations. These findings allow us to propose a model explaining how Hop2-Mnd1 juxtaposes Dmc1-bound ssDNA with distorted recipient double-stranded DNA and thus facilitates strand invasion.
- Department of Biological Sciences, KAIST Institute for the Biocentury, Cancer Metastasis Control Center, Korea Advanced Institute of Science and Technology, Daejeon 305-701, Korea.
Organizational Affiliation: