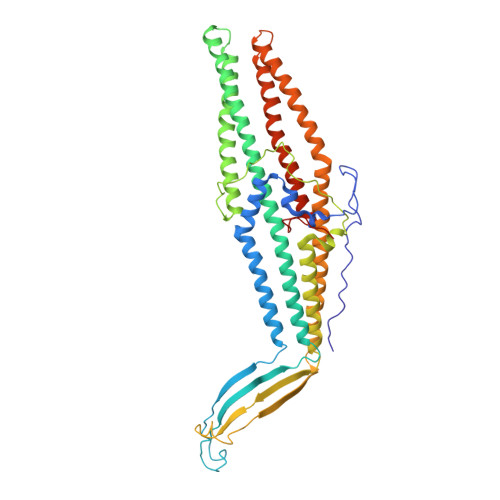
New OprM structure highlighting the nature of the N-terminal anchor.
Monlezun, L., Phan, G., Benabdelhak, H., Lascombe, M.B., Enguene, V.Y., Picard, M., Broutin, I.(2015) Front Microbiol 6: 667-667
- PubMed: 26191054 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fmicb.2015.00667
- Primary Citation Related Structures:
4Y1K - PubMed Abstract:
Among the different mechanisms used by bacteria to resist antibiotics, active efflux plays a major role. In Gram-negative bacteria, active efflux is carried out by tripartite efflux pumps that form a macromolecular assembly spanning both membranes of the cellular wall. At the outer membrane level, a well-conserved outer membrane factor (OMF) protein acts as an exit duct, but its sequence varies greatly among different species. The OMFs share a similar tri-dimensional structure that includes a beta-barrel pore domain that stabilizes the channel within the membrane. In addition, OMFs are often subjected to different N-terminal post-translational modifications (PTMs), such as an acylation with a lipid. The role of additional N-terminal anchors is all the more intriguing since it is not always required among the OMFs family. Understanding this optional PTM could open new research lines in the field of antibiotics resistance. In Escherichia coli, it has been shown that CusC is modified with a tri-acylated lipid, whereas TolC does not show any modification. In the case of OprM from Pseudomonas aeruginosa, the N-terminal modification remains a matter of debate, therefore, we used several approaches to investigate this issue. As definitive evidence, we present a new X-ray structure at 3.8 Å resolution that was solved in a new space group, making it possible to model the N-terminal residue as a palmitoylated cysteine.
- Laboratoire de Cristallographie et RMN Biologiques, CNRS UMR 8015, Faculté de Pharmacie, Université Paris Descartes Paris, France.
Organizational Affiliation: