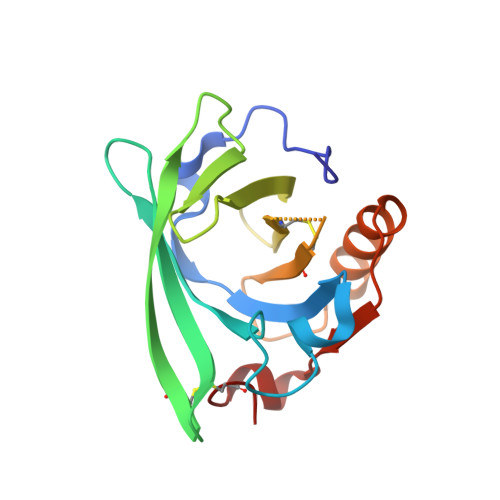
beta-Lactoglobulin interactions with local anaesthetic drugs - Crystallographic and calorimetric studies.
Loch, J.I., Bonarek, P., Polit, A., Jabonski, M., Czub, M., Ye, X., Lewinski, K.(2015) Int J Biol Macromol 80: 87-94
- PubMed: 26092174 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2015.06.013
- Primary Citation Related Structures:
4Y0P, 4Y0Q, 4Y0R, 4Y0S - PubMed Abstract:
Interactions between bovine and goat β-lactoglobulin and tetracaine and pramocaine were investigated with isothermal titration calorimetry, X-ray crystallography and molecular modelling. Tetracaine and pramocaine binding to lactoglobulin is an entropy driven endothermic reaction. In this work, we found that determined association constants and thermodynamic parameters indicate that pramocaine has a higher affinity to lactoglobulin than tetracaine. Crystal structures that were determined with resolutions in the range from 1.90 to 2.30 Å revealed in each case the presence of a single drug molecule bound in the β-barrel in a mode similar to that observed for 14- and 16-carbon fatty acids. The position of the ligand in the β-barrel indicates the optimal fit of 6-carbon aromatic rings to the binding pocket and the major role of hydrophobic interactions in ligand binding. Calculations of tetracaine and pramocaine docking to lactoglobulin revealed that molecular modelling overestimated the role of polar protein-drug interactions.
- Department of Crystal Chemistry and Crystal Physics, Faculty of Chemistry, Jagiellonian University in Kraków, Ingardena 3, 30-060 Kraków, Poland.
Organizational Affiliation: