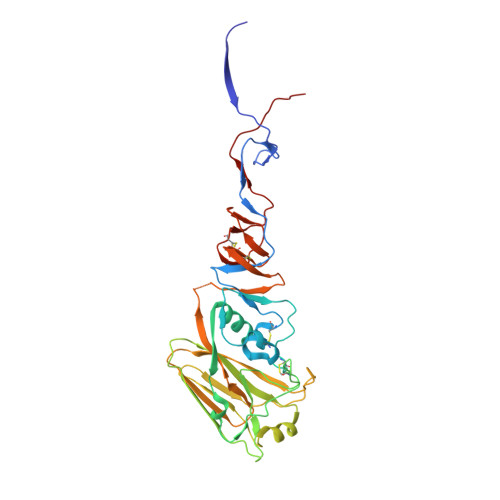
Structure and Receptor Binding of the Hemagglutinin from a Human H6N1 Influenza Virus.
Tzarum, N., de Vries, R.P., Zhu, X., Yu, W., McBride, R., Paulson, J.C., Wilson, I.A.(2015) Cell Host Microbe 17: 369-376
- PubMed: 25766295
- DOI: https://doi.org/10.1016/j.chom.2015.02.005
- Primary Citation Related Structures:
4XKD, 4XKE, 4XKF, 4XKG - PubMed Abstract:
Avian influenza viruses that cause infection and are transmissible in humans involve changes in the receptor binding site (RBS) of the viral hemagglutinin (HA) that alter receptor preference from α2-3-linked (avian-like) to α2-6-linked (human-like) sialosides. A human case of avian-origin H6N1 influenza virus was recently reported, but the molecular mechanisms contributing to it crossing the species barrier are unknown. We find that, although the H6 HA RBS contains D190V and G228S substitutions that potentially promote human receptor binding, recombinant H6 HA preferentially binds α2-3-linked sialosides, indicating no adaptation to human receptors. Crystal structures of H6 HA with avian and human receptor analogs reveal that H6 HA preferentially interacts with avian receptor analogs. This binding mechanism differs from other HA subtypes due to a unique combination of RBS residues, highlighting additional variation in HA-receptor interactions and the challenges in predicting which influenza strains and subtypes can infect humans and cause pandemics.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: