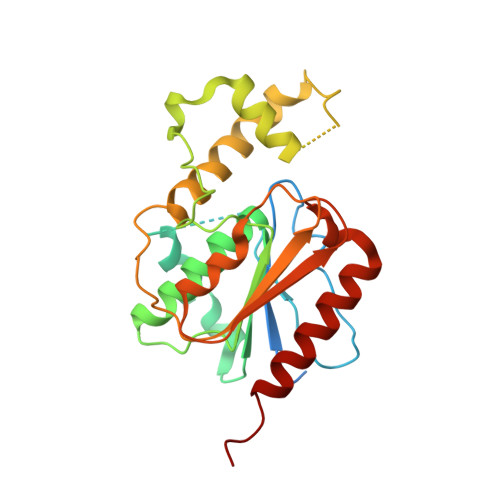
Crystal Structure and Substrate Specificity of Human Thioesterase 2: INSIGHTS INTO THE MOLECULAR BASIS FOR THE MODULATION OF FATTY ACID SYNTHASE.
Ritchie, M.K., Johnson, L.C., Clodfelter, J.E., Pemble, C.W., Fulp, B.E., Furdui, C.M., Kridel, S.J., Lowther, W.T.(2016) J Biological Chem 291: 3520-3530
- PubMed: 26663084 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.702597
- Primary Citation Related Structures:
4XJV - PubMed Abstract:
The type I fatty acid synthase (FASN) is responsible for the de novo synthesis of palmitate. Chain length selection and release is performed by the C-terminal thioesterase domain (TE1). FASN expression is up-regulated in cancer, and its activity levels are controlled by gene dosage and transcriptional and post-translational mechanisms. In addition, the chain length of fatty acids produced by FASN is controlled by a type II thioesterase called TE2 (E.C. 3.1.2.14). TE2 has been implicated in breast cancer and generates a broad lipid distribution within milk. The molecular basis for the ability of the TE2 to compete with TE1 for the acyl chain attached to the acyl carrier protein (ACP) domain of FASN is unknown. Herein, we show that human TE1 efficiently hydrolyzes acyl-CoA substrate mimetics. In contrast, TE2 prefers an engineered human acyl-ACP substrate and readily releases short chain fatty acids from full-length FASN during turnover. The 2.8 Å crystal structure of TE2 reveals a novel capping domain insert within the α/β hydrolase core. This domain is reminiscent of capping domains of type II thioesterases involved in polyketide synthesis. The structure also reveals that the capping domain had collapsed onto the active site containing the Ser-101-His-237-Asp-212 catalytic triad. This observation suggests that the capping domain opens to enable the ACP domain to dock and to place the acyl chain and 4'-phosphopantetheinyl-linker arm correctly for catalysis. Thus, the ability of TE2 to prematurely release fatty acids from FASN parallels the role of editing thioesterases involved in polyketide and non-ribosomal peptide synthase synthases.
- From the Center for Structural Biology and Department of Biochemistry.
Organizational Affiliation: