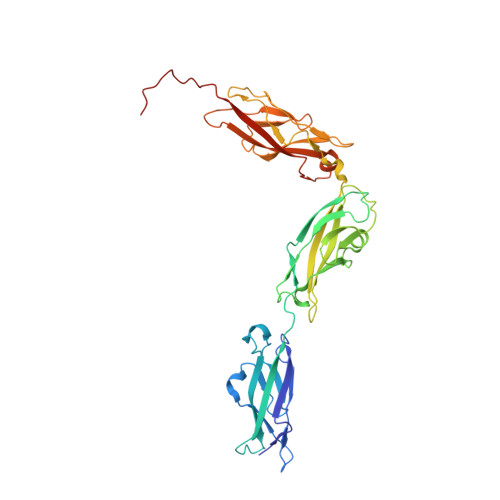
An elastic element in the protocadherin-15 tip link of the inner ear.
Araya-Secchi, R., Neel, B.L., Sotomayor, M.(2016) Nat Commun 7: 13458-13458
- PubMed: 27857071 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms13458
- Primary Citation Related Structures:
4XHZ, 5KJ4 - PubMed Abstract:
Tip link filaments convey force and gate inner-ear hair-cell transduction channels to mediate perception of sound and head movements. Cadherin-23 and protocadherin-15 form tip links through a calcium-dependent interaction of their extracellular domains made of multiple extracellular cadherin (EC) repeats. These repeats are structurally similar, but not identical in sequence, often featuring linkers with conserved calcium-binding sites that confer mechanical strength to them. Here we present the X-ray crystal structures of human protocadherin-15 EC8-EC10 and mouse EC9-EC10, which show an EC8-9 canonical-like calcium-binding linker, and an EC9-10 calcium-free linker that alters the linear arrangement of EC repeats. Molecular dynamics simulations and small-angle X-ray scattering experiments support this non-linear conformation. Simulations also suggest that unbending of EC9-10 confers some elasticity to otherwise rigid tip links. The new structure provides a first view of protocadherin-15's non-canonical EC linkers and suggests how they may function in inner-ear mechanotransduction, with implications for other cadherins.
- Department of Chemistry and Biochemistry, The Ohio State University, 484 W 12th Avenue, Columbus, Ohio 43210, USA.
Organizational Affiliation: