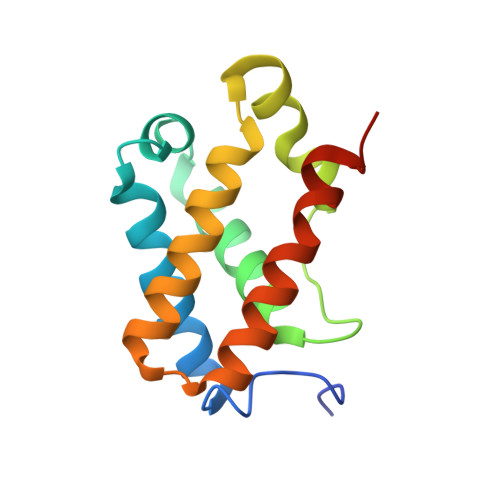
Structure of Chlamydomonas reinhardtii THB1, a group 1 truncated hemoglobin with a rare histidine-lysine heme ligation.
Rice, S.L., Boucher, L.E., Schlessman, J.L., Preimesberger, M.R., Bosch, J., Lecomte, J.T.(2015) Acta Crystallogr F Struct Biol Commun 71: 718-725
- PubMed: 26057801
- DOI: https://doi.org/10.1107/S2053230X15006949
- Primary Citation Related Structures:
4XDI - PubMed Abstract:
THB1 is one of several group 1 truncated hemoglobins (TrHb1s) encoded in the genome of the unicellular green alga Chlamydomonas reinhardtii. THB1 expression is under the control of NIT2, the master regulator of nitrate assimilation, which also controls the expression of the only nitrate reductase in the cell, NIT1. In vitro and physiological evidence suggests that THB1 converts the nitric oxide generated by NIT1 into nitrate. To aid in the elucidation of the function and mechanism of THB1, the structure of the protein was solved in the ferric state. THB1 resembles other TrHb1s, but also exhibits distinct features associated with the coordination of the heme iron by a histidine (proximal) and a lysine (distal). The new structure illustrates the versatility of the TrHb1 fold, suggests factors that stabilize the axial ligation of a lysine, and highlights the difficulty of predicting the identity of the distal ligand, if any, in this group of proteins.
- T. C. Jenkins Department of Biophysics, Johns Hopkins University, 3400 North Charles Street, Baltimore, MD 21218, USA.
Organizational Affiliation: