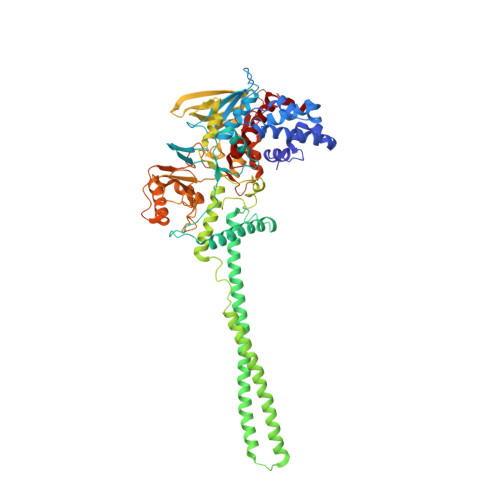
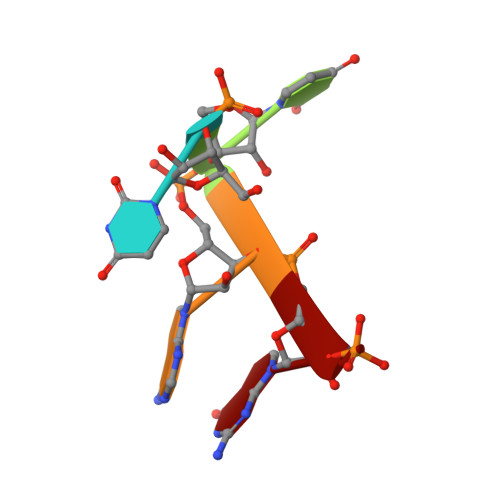
G-quadruplex RNA binding and recognition by the lysine-specific histone demethylase-1 enzyme.
Hirschi, A., Martin, W.J., Luka, Z., Loukachevitch, L.V., Reiter, N.J.(2016) RNA 22: 1250-1260
- PubMed: 27277658 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.057265.116
- Primary Citation Related Structures:
4XBF - PubMed Abstract:
Lysine-specific histone demethylase 1 (LSD1) is an essential epigenetic regulator in metazoans and requires the co-repressor element-1 silencing transcription factor (CoREST) to efficiently catalyze the removal of mono- and dimethyl functional groups from histone 3 at lysine positions 4 and 9 (H3K4/9). LSD1 interacts with over 60 regulatory proteins and also associates with lncRNAs (TERRA, HOTAIR), suggesting a regulatory role for RNA in LSD1 function. We report that a stacked, intramolecular G-quadruplex (GQ) forming TERRA RNA (GG[UUAGGG]8UUA) binds tightly to the functional LSD1-CoREST complex (Kd ≈ 96 nM), in contrast to a single GQ RNA unit ([UUAGGG]4U), a GQ DNA ([TTAGGG]4T), or an unstructured single-stranded RNA. Stabilization of a parallel-stranded GQ RNA structure by monovalent potassium ions (K(+)) is required for high affinity binding to the LSD1-CoREST complex. These data indicate that LSD1 can distinguish between RNA and DNA as well as structured versus unstructured nucleotide motifs. Further, cross-linking mass spectrometry identified the primary location of GQ RNA binding within the SWIRM/amine oxidase domain (AOD) of LSD1. An ssRNA binding region adjacent to this GQ binding site was also identified via X-ray crystallography. This RNA binding interface is consistent with kinetic assays, demonstrating that a GQ-forming RNA can serve as a noncompetitive inhibitor of LSD1-catalyzed demethylation. The identification of a GQ RNA binding site coupled with kinetic data suggests that structured RNAs can function as regulatory molecules in LSD1-mediated mechanisms.
- Department of Biochemistry, Vanderbilt University Medical Center, Nashville, Tennessee 37232-0146, USA.
Organizational Affiliation: