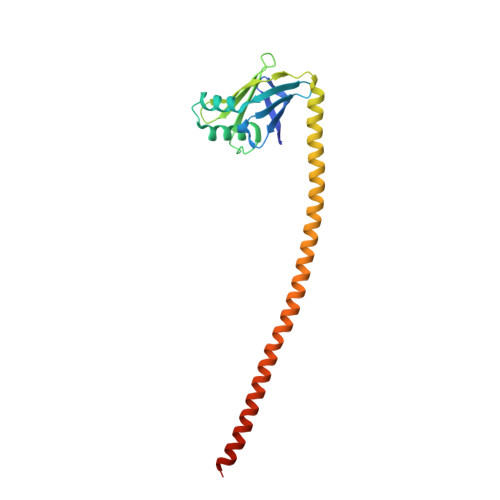
Skip residues modulate the structural properties of the myosin rod and guide thick filament assembly.
Taylor, K.C., Buvoli, M., Korkmaz, E.N., Buvoli, A., Zheng, Y., Heinze, N.T., Cui, Q., Leinwand, L.A., Rayment, I.(2015) Proc Natl Acad Sci U S A 112: E3806-E3815
- PubMed: 26150528 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1505813112
- Primary Citation Related Structures:
4XA1, 4XA3, 4XA4, 4XA6 - PubMed Abstract:
The rod of sarcomeric myosins directs thick filament assembly and is characterized by the insertion of four skip residues that introduce discontinuities in the coiled-coil heptad repeats. We report here that the regions surrounding the first three skip residues share high structural similarity despite their low sequence homology. Near each of these skip residues, the coiled-coil transitions to a nonclose-packed structure inducing local relaxation of the superhelical pitch. Moreover, molecular dynamics suggest that these distorted regions can assume different conformationally stable states. In contrast, the last skip residue region constitutes a true molecular hinge, providing C-terminal rod flexibility. Assembly of myosin with mutated skip residues in cardiomyocytes shows that the functional importance of each skip residue is associated with rod position and reveals the unique role of the molecular hinge in promoting myosin antiparallel packing. By defining the biophysical properties of the rod, the structures and molecular dynamic calculations presented here provide insight into thick filament formation, and highlight the structural differences occurring between the coiled-coils of myosin and the stereotypical tropomyosin. In addition to extending our knowledge into the conformational and biological properties of coiled-coil discontinuities, the molecular characterization of the four myosin skip residues also provides a guide to modeling the effects of rod mutations causing cardiac and skeletal myopathies.
- Department of Biochemistry, University of Wisconsin, Madison, WI 53706;
Organizational Affiliation: