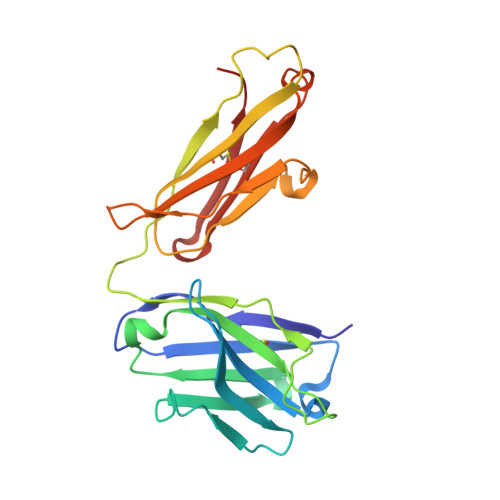
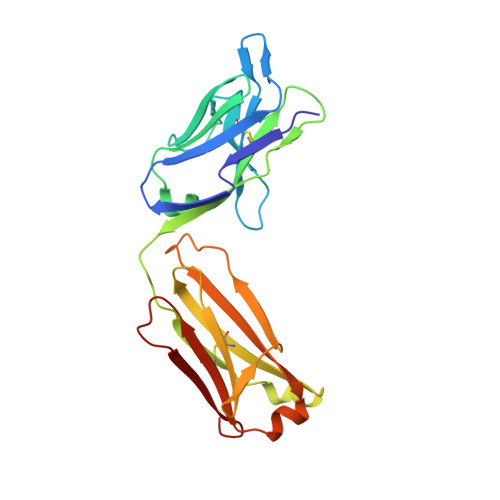
Structure of the omalizumab Fab
Jensen, R.K., Plum, M., Tjerrild, L., Jakob, T., Spillner, E., Andersen, G.R.(2015) Acta Crystallogr F Struct Biol Commun 71: 419-426
- PubMed: 25849503 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X15004100
- Primary Citation Related Structures:
4X7S, 4X7T - PubMed Abstract:
Omalizumab is a humanized anti-IgE antibody that inhibits the binding of IgE to its receptors on mast cells and basophils, thus blocking the IgE-mediated release of inflammatory mediators from these cells. Omalizumab binds to the Fc domains of IgE in proximity to the binding site of the high-affinity IgE receptor FcℇRI, but the epitope and the mechanisms and conformations governing the recognition remain unknown. In order to elucidate the molecular mechanism of its anti-IgE activity, the aim was to analyse the interaction of omalizumab with human IgE. Therefore, IgE Fc Cℇ2-4 was recombinantly produced in mammalian HEK-293 cells. Functionality of the IgE Fc was proven by ELISA and mediator-release assays. Omalizumab IgG was cleaved with papain and the resulting Fab was purified by ion-exchange chromatography. The complex of IgE Fc with omalizumab was prepared by size-exclusion chromatography. However, crystals containing the complex were not obtained, suggesting that the process of crystallization favoured the dissociation of the two proteins. Instead, two structures of the omalizumab Fab with maximum resolutions of 1.9 and 3.0 Å were obtained. The structures reveal the arrangement of the CDRs and the position of omalizumab residues known from prior functional studies to be involved in IgE binding. Thus, the structure of omalizumab provides the structural basis for understanding the function of omalizumab, allows optimization of the procedure for complex crystallization and poses questions about the conformational requirements for anti-IgE activity.
- Department of Molecular Biology and Genetics, Aarhus University, Gustav Wiedsvej 10C, 8000 Aarhus, Denmark.
Organizational Affiliation: