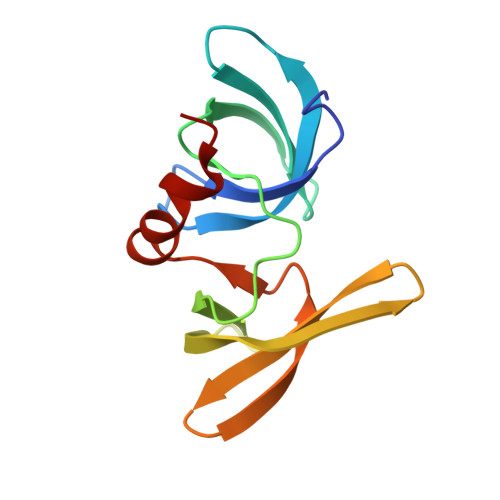
An Acetyl-Methyl Switch Drives a Conformational Change in p53.
Tong, Q., Mazur, S.J., Rincon-Arano, H., Rothbart, S.B., Kuznetsov, D.M., Cui, G., Liu, W.H., Gete, Y., Klein, B.J., Jenkins, L., Mer, G., Kutateladze, A.G., Strahl, B.D., Groudine, M., Appella, E., Kutateladze, T.G.(2015) Structure 23: 322-331
- PubMed: 25651062 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2014.12.010
- Primary Citation Related Structures:
4X34 - PubMed Abstract:
Individual posttranslational modifications (PTMs) of p53 mediate diverse p53-dependent responses; however, much less is known about the combinatorial action of adjacent modifications. Here, we describe crosstalk between the early DNA damage response mark p53K382me2 and the surrounding PTMs that modulate binding of p53 cofactors, including 53BP1 and p300. The 1.8 Å resolution crystal structure of the tandem Tudor domain (TTD) of 53BP1 in complex with p53 peptide acetylated at K381 and dimethylated at K382 (p53K381acK382me2) reveals that the dual PTM induces a conformational change in p53. The α-helical fold of p53K381acK382me2 positions the side chains of R379, K381ac, and K382me2 to interact with TTD concurrently, reinforcing a modular design of double PTM mimetics. Biochemical and nuclear magnetic resonance analyses show that other surrounding PTMs, including phosphorylation of serine/threonine residues of p53, affect association with TTD. Our findings suggest a novel PTM-driven conformation switch-like mechanism that may regulate p53 interactions with binding partners.
- Department of Pharmacology, University of Colorado School of Medicine, Aurora, CO 80045, USA.
Organizational Affiliation: