Crystal structure of acetylcholine binding protein (AChBP) from Aplysia Californica in complex with click chemistry compound.
Talley, T.T.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Soluble acetylcholine receptor | 216 | Aplysia californica | Mutation(s): 0 | 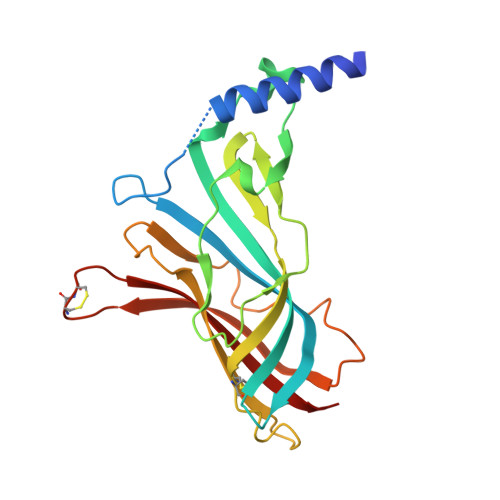 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q8WSF8 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| MD4 Download:Ideal Coordinates CCD File | F [auth A], G [auth D] | (3-exo)-8,8-dimethyl-3-[4-(pyridin-4-yl)-1H-1,2,3-triazol-1-yl]-8-azoniabicyclo[3.2.1]octane C16 H22 N5 IRDLJZVYUZKXLE-QKDCVEJESA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 87.014 | α = 90 |
| b = 115.391 | β = 90 |
| c = 130.26 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data reduction |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of Neurological Disorders and Stroke (NIH/NINDS) | United States | F32 NS043063-03 |