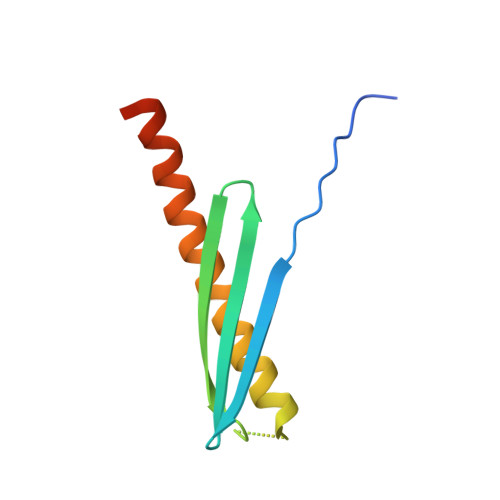
A functional role of Rv1738 in Mycobacterium tuberculosis persistence suggested by racemic protein crystallography.
Bunker, R.D., Mandal, K., Bashiri, G., Chaston, J.J., Pentelute, B.L., Lott, J.S., Kent, S.B., Baker, E.N.(2015) Proc Natl Acad Sci U S A 112: 4310-4315
- PubMed: 25831534 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1422387112
- Primary Citation Related Structures:
4WPY, 4WSP - PubMed Abstract:
Protein 3D structure can be a powerful predictor of function, but it often faces a critical roadblock at the crystallization step. Rv1738, a protein from Mycobacterium tuberculosis that is strongly implicated in the onset of nonreplicating persistence, and thereby latent tuberculosis, resisted extensive attempts at crystallization. Chemical synthesis of the L- and D-enantiomeric forms of Rv1738 enabled facile crystallization of the D/L-racemic mixture. The structure was solved by an ab initio approach that took advantage of the quantized phases characteristic of diffraction by centrosymmetric crystals. The structure, containing L- and D-dimers in a centrosymmetric space group, revealed unexpected homology with bacterial hibernation-promoting factors that bind to ribosomes and suppress translation. This suggests that the functional role of Rv1738 is to contribute to the shutdown of ribosomal protein synthesis during the onset of nonreplicating persistence of M. tuberculosis.
- Maurice Wilkins Centre for Molecular Biodiscovery and School of Biological Sciences, University of Auckland, Auckland 1142, New Zealand; and.
Organizational Affiliation: