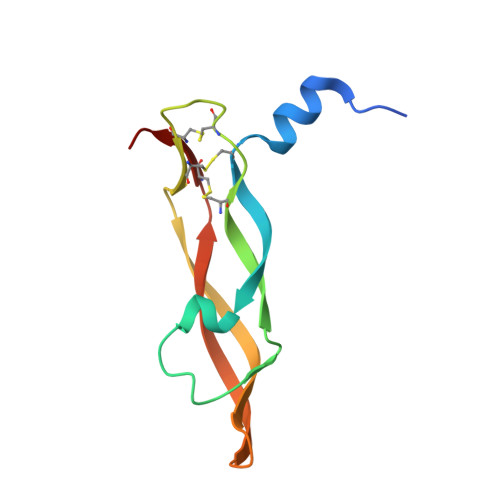
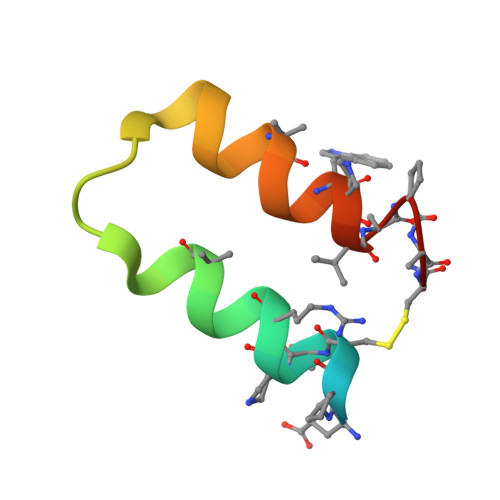
Targeting diverse protein-protein interaction interfaces with alpha / beta-peptides derived from the Z-domain scaffold.
Checco, J.W., Kreitler, D.F., Thomas, N.C., Belair, D.G., Rettko, N.J., Murphy, W.L., Forest, K.T., Gellman, S.H.(2015) Proc Natl Acad Sci U S A 112: 4552-4557
- PubMed: 25825775 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1420380112
- Primary Citation Related Structures:
4WPB - PubMed Abstract:
Peptide-based agents derived from well-defined scaffolds offer an alternative to antibodies for selective and high-affinity recognition of large and topologically complex protein surfaces. Here, we describe a strategy for designing oligomers containing both α- and β-amino acid residues ("α/β-peptides") that mimic several peptides derived from the three-helix bundle "Z-domain" scaffold. We show that α/β-peptides derived from a Z-domain peptide targeting vascular endothelial growth factor (VEGF) can structurally and functionally mimic the binding surface of the parent peptide while exhibiting significantly decreased susceptibility to proteolysis. The tightest VEGF-binding α/β-peptide inhibits the VEGF165-induced proliferation of human umbilical vein endothelial cells. We demonstrate the versatility of this strategy by showing how principles underlying VEGF signaling inhibitors can be rapidly extended to produce Z-domain-mimetic α/β-peptides that bind to two other protein partners, IgG and tumor necrosis factor-α. Because well-established selection techniques can identify high-affinity Z-domain derivatives from large DNA-encoded libraries, our findings should enable the design of biostable α/β-peptides that bind tightly and specifically to diverse targets of biomedical interest. Such reagents would be useful for diagnostic and therapeutic applications.
- Department of Chemistry, University of Wisconsin-Madison, Madison, WI 53706;
Organizational Affiliation: