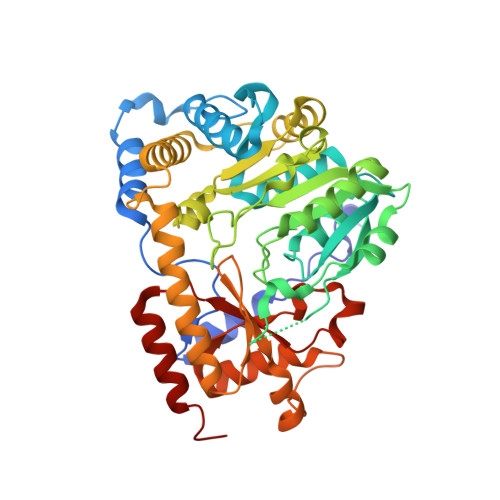
High resolution crystal structures of human kynurenine aminotransferase-I bound to PLP cofactor, and in complex with aminooxyacetate.
Nadvi, N.A., Salam, N.K., Park, J., Akladios, F.N., Kapoor, V., Collyer, C.A., Gorrell, M.D., Church, W.B.(2017) Protein Sci 26: 727-736
- PubMed: 28097769 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3119
- Primary Citation Related Structures:
4WLH, 4WLJ - PubMed Abstract:
In this study, we report two high-resolution structures of the pyridoxal 5' phosphate (PLP)-dependent enzyme kynurenine aminotransferase-I (KAT-I). One is the native structure with the cofactor in the PLP form bound to Lys247 with the highest resolution yet available for KAT-I at 1.28 Å resolution, and the other with the general PLP-dependent aminotransferase inhibitor, aminooxyacetate (AOAA) covalently bound to the cofactor at 1.54 Å. Only small conformational differences are observed in the vicinity of the aldimine (oxime) linkage with which the PLP forms the Schiff base with Lys247 in the 1.28 Å resolution native structure, in comparison to other native PLP-bound structures. We also report the inhibition of KAT-1 by AOAA and aminooxy-phenylpropionic acid (AOPP), with IC50s of 13.1 and 5.7 μM, respectively. The crystal structure of the enzyme in complex with the inhibitor AOAA revealed that the cofactor is the PLP form with the external aldimine linkage. The location of this oxime with the PLP, which forms in place of the native internal aldimine linkage of PLP of the native KAT-I, is away from the position of the native internal aldimine, with the free Lys247 substantially retaining the orientation of the native structure. Tyr101, at the active site, was observed in two conformations in both structures.
- Group in Biomolecular Structure and Informatics, Faculty of Pharmacy, University of Sydney, Sydney, New South Wales, Australia.
Organizational Affiliation: