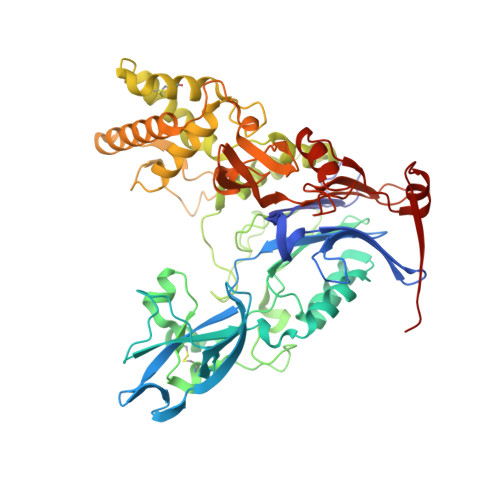
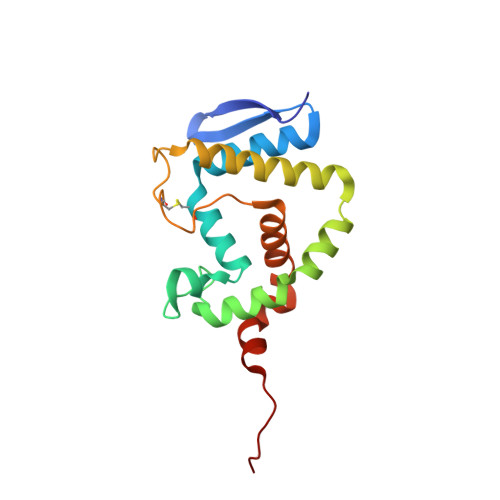
n-Alkylboronic Acid Inhibitors Reveal Determinants of Ligand Specificity in the Quorum-Quenching and Siderophore Biosynthetic Enzyme PvdQ.
Clevenger, K.D., Wu, R., Liu, D., Fast, W.(2014) Biochemistry 53: 6679-6686
- PubMed: 25290020 Search on PubMed
- DOI: https://doi.org/10.1021/bi501086s
- Primary Citation Related Structures:
4WKS, 4WKT, 4WKU, 4WKV - PubMed Abstract:
The enzyme PvdQ (E.C. 3.5.1.97) from Pseudomonas aeruginosa is an N-terminal nucleophile hydrolase that catalyzes the removal of an N-myristyl substituent from a biosynthetic precursor of the iron-chelating siderophore pyoverdine. Inhibitors of pyoverdine biosynthesis are potential antibiotics since iron is essential for growth and scarce in most infections. PvdQ also catalyzes hydrolytic amide bond cleavage of selected N-acyl-l-homoserine lactone quorum-sensing signals used by some Gram-negative pathogens to coordinate the transcription of virulence factors. The resulting quorum-quenching activity of PvdQ has potential applications in antivirulence therapies. To inform both inhibitor design and enzyme engineering efforts, a series of n-alkylboronic acid inhibitors of PvdQ was characterized to reveal determinants of ligand selectivity. A simple homologation series results in compounds with Ki values that span from 4.7 mM to 190 pM, with a dependence of ΔGbind values on chain length of -1.0 kcal/mol/CH2. X-ray crystal structures are determined for the PvdQ complexes with 1-ethyl-, 1-butyl-, 1-hexyl-, and 1-octylboronic acids at 1.6, 1.8, 2.0, and 2.1 Å resolution, respectively. The 1-hexyl- and 1-octylboronic acids form tetrahedral adducts with the active-site N-terminal Ser217 in the β-subunit of PvdQ, and the n-alkyl substituents are bound in the acyl-group binding site. The 1-ethyl- and 1-butylboronic acids also form adducts with Ser217 but instead form trigonal planar adducts and extend their n-alkyl substituents into an alternative binding site. These results are interpreted to propose a ligand discrimination model for PvdQ that informs the development of PvdQ-related tools and therapeutics.
- Medicinal Chemistry Division, College of Pharmacy, †Biochemistry Graduate Program, and ⊥Center for Infectious Disease, The University of Texas , Austin, Texas 78712, United States.
Organizational Affiliation: