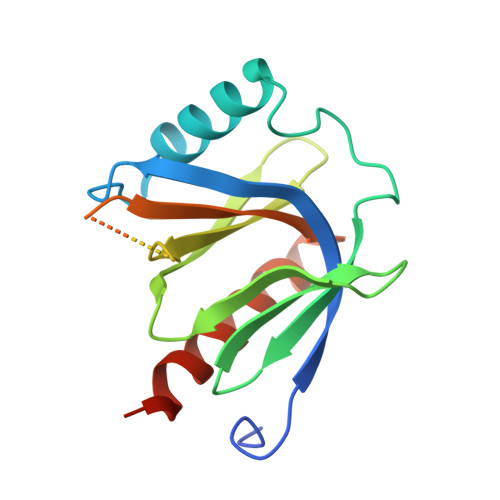
Structural Basis for the Disruption of the Cerebral Cavernous Malformations 2 (CCM2) Interaction with Krev Interaction Trapped 1 (KRIT1) by Disease-associated Mutations.
Fisher, O.S., Liu, W., Zhang, R., Stiegler, A.L., Ghedia, S., Weber, J.L., Boggon, T.J.(2015) J Biological Chem 290: 2842-2853
- PubMed: 25525273 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.616433
- Primary Citation Related Structures:
4WJ7 - PubMed Abstract:
Familial cerebral cavernous malformations (CCMs) are predominantly neurovascular lesions and are associated with mutations within the KRIT1, CCM2, and PDCD10 genes. The protein products of KRIT1 and CCM2 (Krev interaction trapped 1 (KRIT1) and cerebral cavernous malformations 2 (CCM2), respectively) directly interact with each other. Disease-associated mutations in KRIT1 and CCM2 mostly result in loss of their protein products, although rare missense point mutations can also occur. From gene sequencing of patients known or suspected to have one or more CCMs, we discover a series of missense point mutations in KRIT1 and CCM2 that result in missense mutations in the CCM2 and KRIT1 proteins. To place these mutations in the context of the molecular level interactions of CCM2 and KRIT1, we map the interaction of KRIT1 and CCM2 and find that the CCM2 phosphotyrosine binding (PTB) domain displays a preference toward the third of the three KRIT1 NPX(Y/F) motifs. We determine the 2.75 Å co-crystal structure of the CCM2 PTB domain with a peptide corresponding to KRIT1(NPX(Y/F)3), revealing a Dab-like PTB fold for CCM2 and its interaction with KRIT1(NPX(Y/F)3). We find that several disease-associated missense mutations in CCM2 have the potential to interrupt the KRIT1-CCM2 interaction by destabilizing the CCM2 PTB domain and that a KRIT1 mutation also disrupts this interaction. We therefore provide new insights into the architecture of CCM2 and how the CCM complex is disrupted in CCM disease.
- From the Department of Pharmacology, Yale University School of Medicine, New Haven, Connecticut 06520.
Organizational Affiliation: