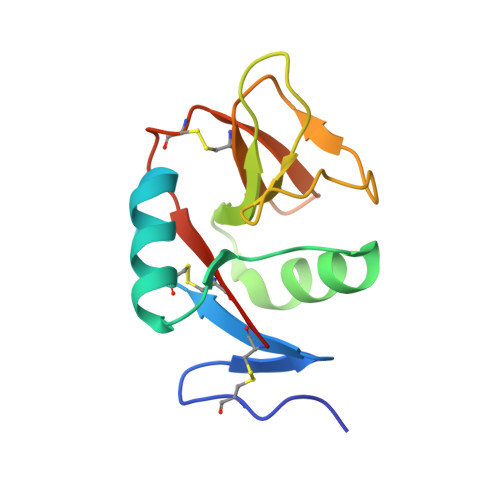
Crystal structure of extracellular domain of human lectin-like transcript 1 (LLT1), the ligand for natural killer receptor-P1A
Kita, S., Matsubara, H., Kasai, Y., Tamaoki, T., Okabe, Y., Fukuhara, H., Kamishikiryo, J., Krayukhina, E., Uchiyama, S., Ose, T., Kuroki, K., Maenaka, K.(2015) Eur J Immunol 45: 1605-1613
- PubMed: 25826155 Search on PubMed
- DOI: https://doi.org/10.1002/eji.201545509
- Primary Citation Related Structures:
4WCO - PubMed Abstract:
Emerging evidence has revealed the pivotal roles of C-type lectin-like receptors (CTLRs) in the regulation of a wide range of immune responses. Human natural killer cell receptor-P1A (NKRP1A) is one of the CTLRs and recognizes another CTLR, lectin-like transcript 1 (LLT1) on target cells to control NK, NKT and Th17 cells. The structural basis for the NKRP1A-LLT1 interaction was limitedly understood. Here, we report the crystal structure of the ectodomain of LLT1. The plausible receptor-binding face of the C-type lectin-like domain is flat, and forms an extended β-sheet. The residues of this face are relatively conserved with another CTLR, keratinocyte-associated C-type lectin, which binds to the CTLR member, NKp65. A LLT1-NKRP1A complex model, prepared using the crystal structures of LLT1 and the keratinocyte-associated C-type lectin-NKp65 complex, reasonably satisfies the charge consistency and the conformational complementarity to explain a previous mutagenesis study. Furthermore, crystal packing and analytical ultracentrifugation revealed dimer formation, which supports a complex model. Our results provide structural insights for understanding the binding modes and signal transduction mechanisms, which are likely to be conserved in the CTLR family, and for further rational drug design towards regulating the LLT1 function.
- Laboratory of Biomolecular Science, Faculty of Pharmaceutical Sciences, Hokkaido University, Sapporo, Japan.
Organizational Affiliation: