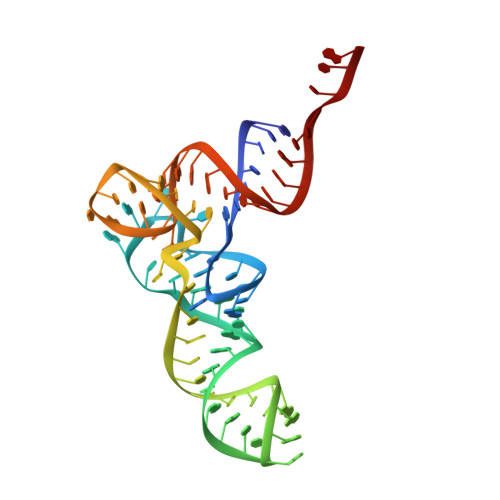
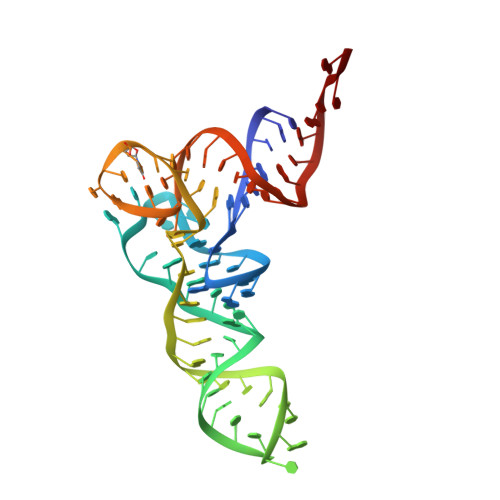
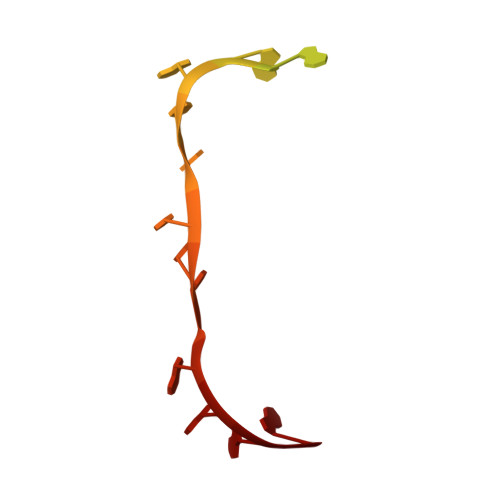
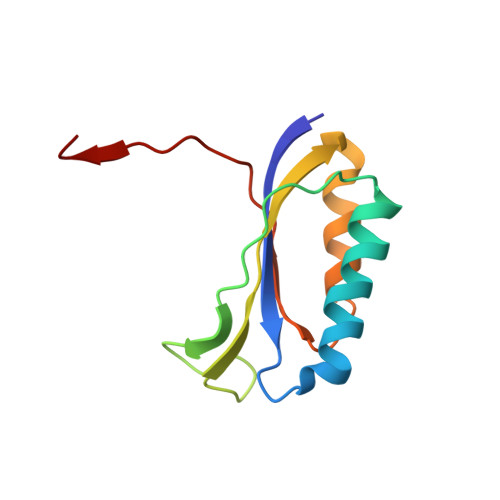
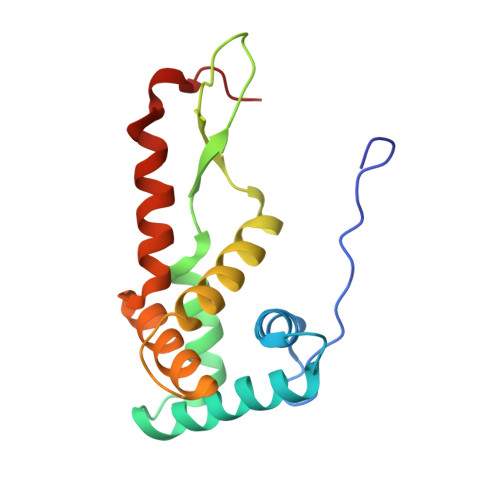
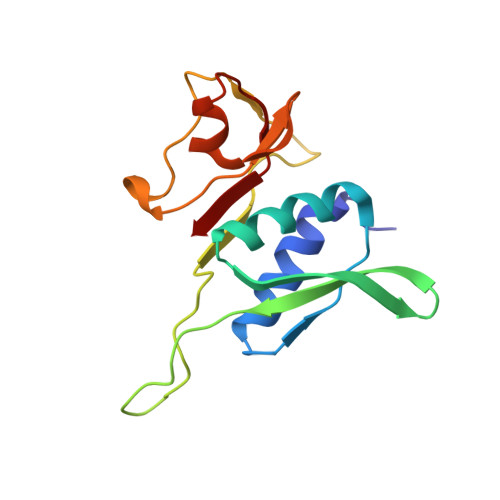
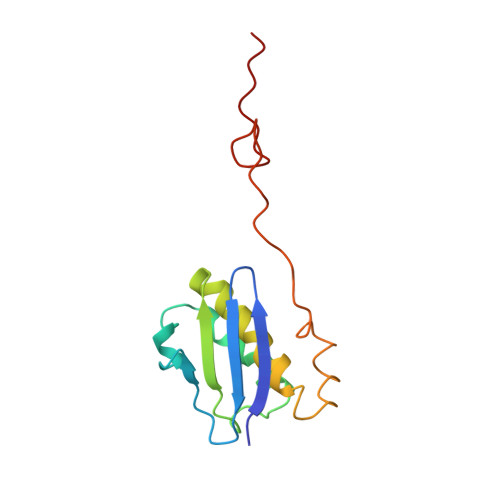
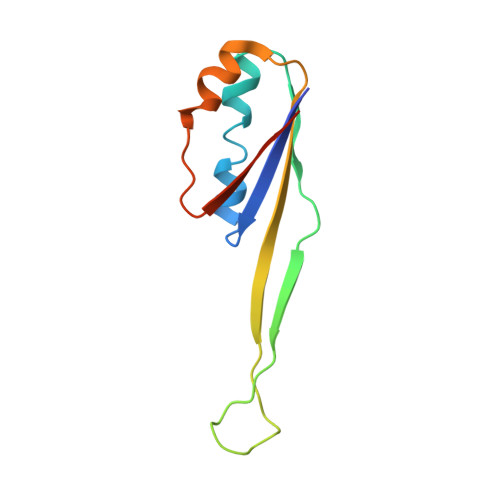
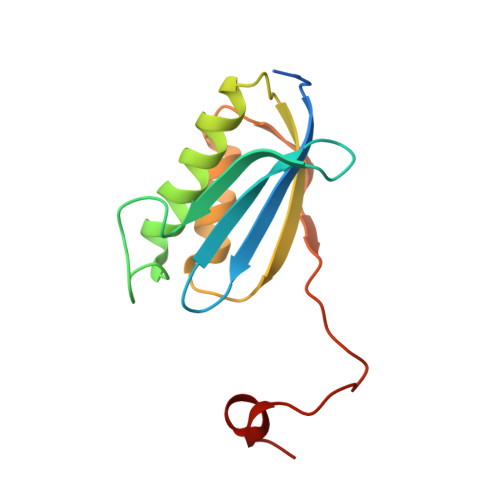
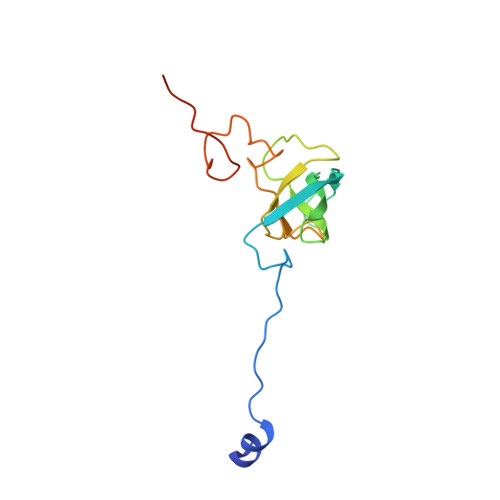
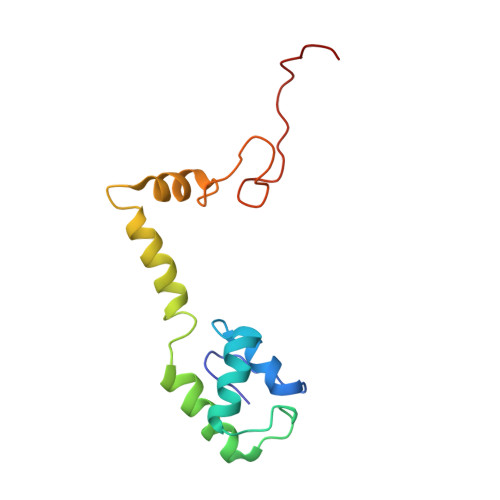
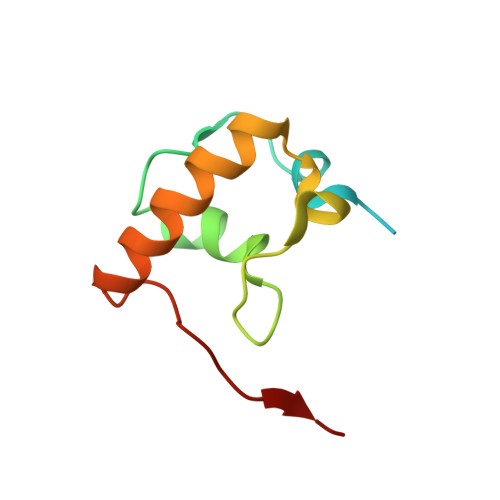
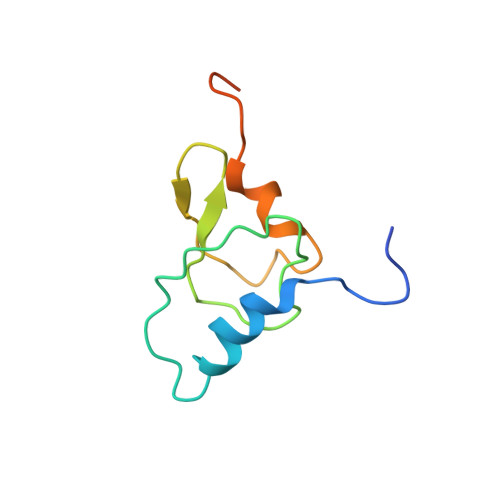
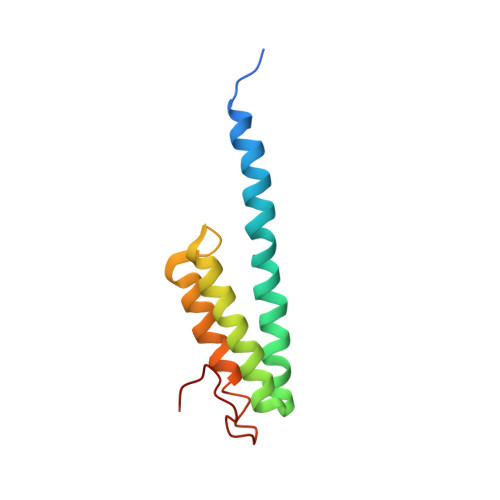
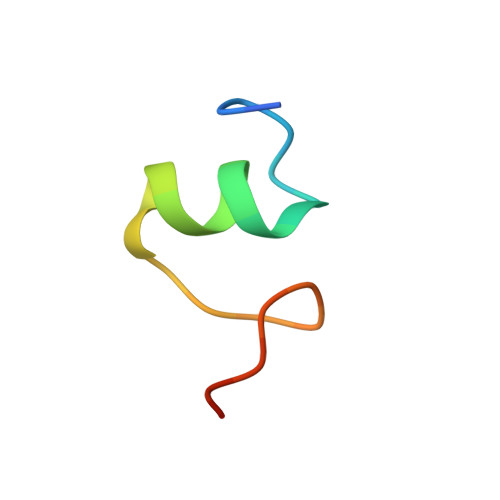
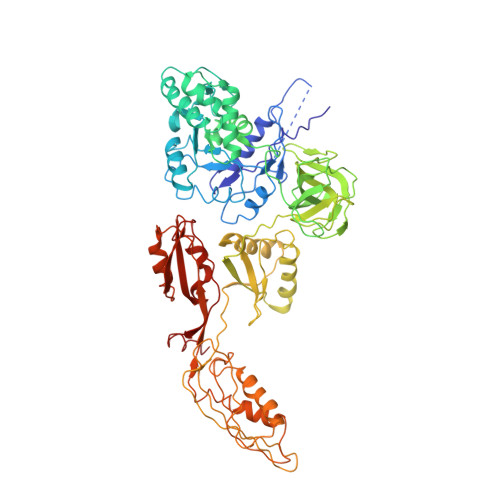
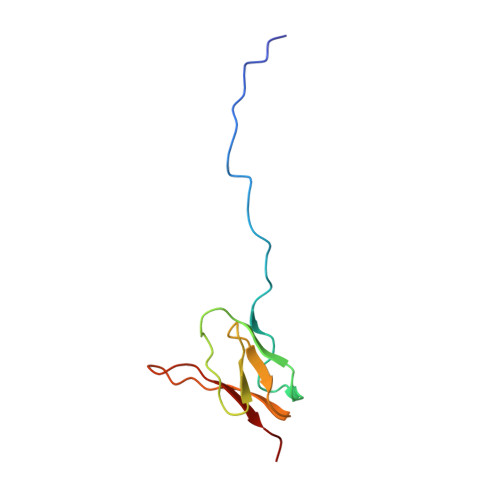
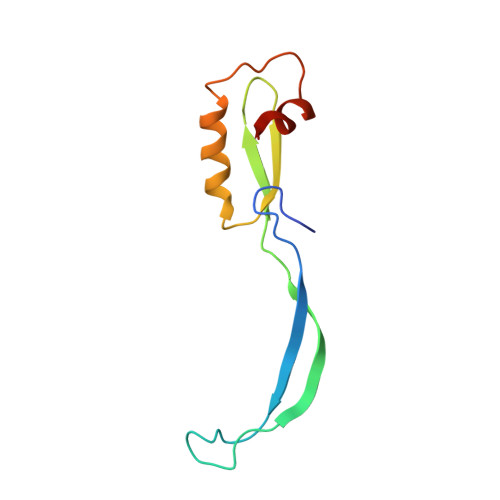
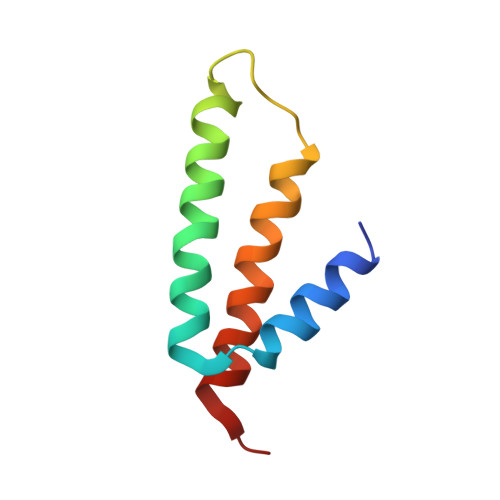
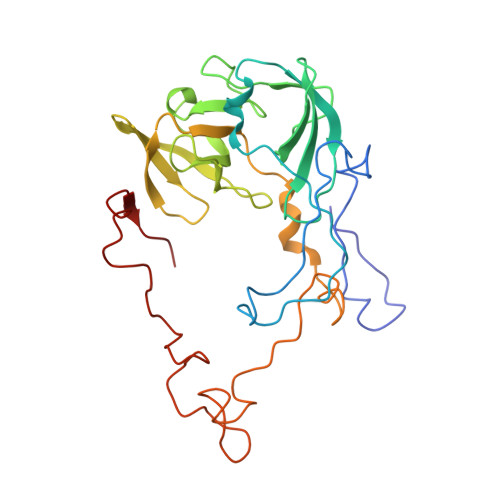
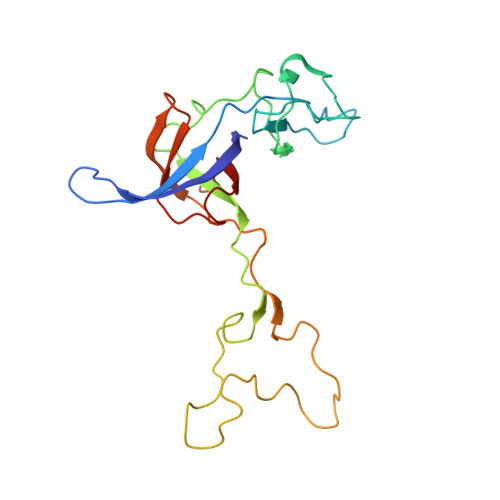
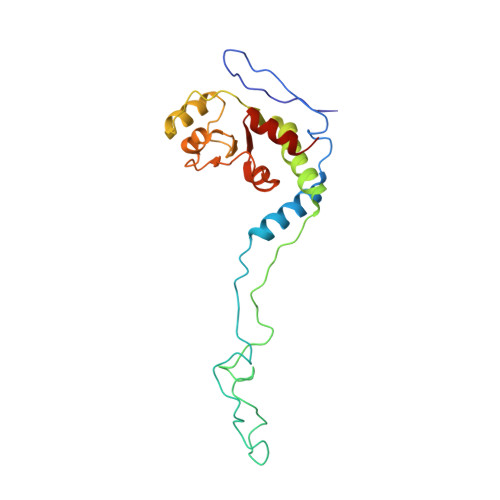
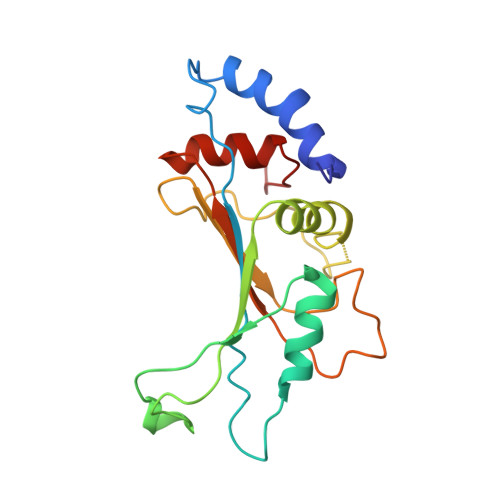
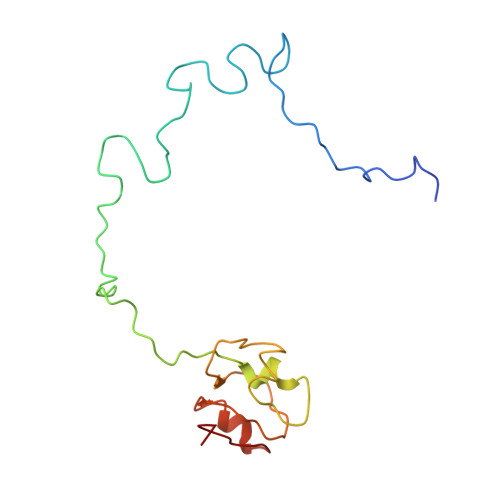
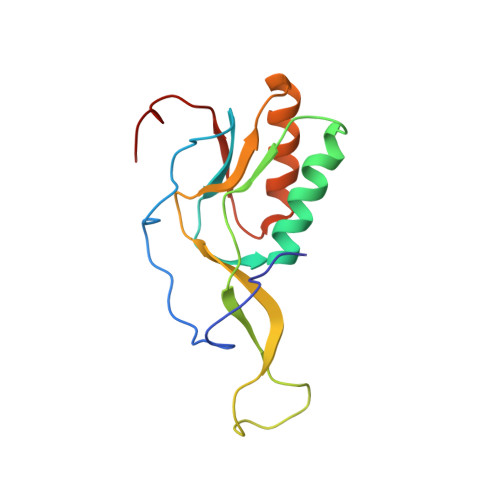
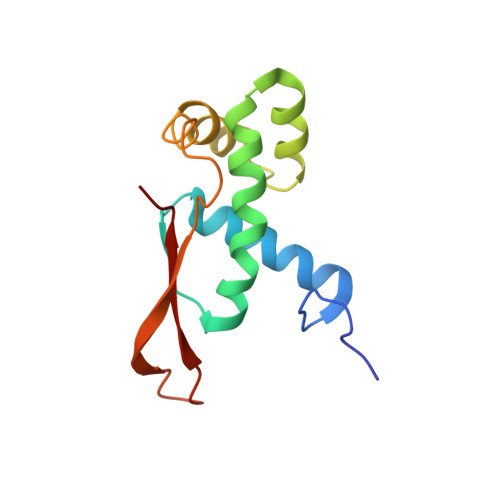
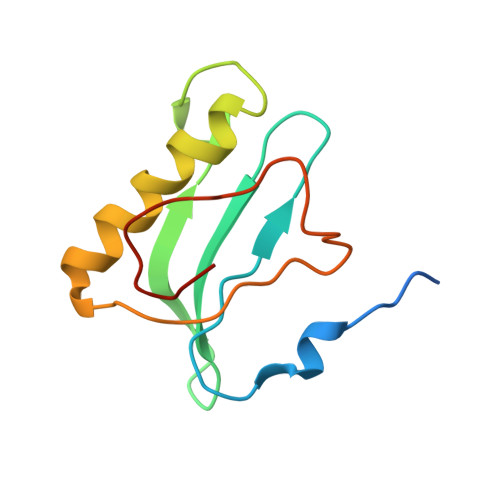
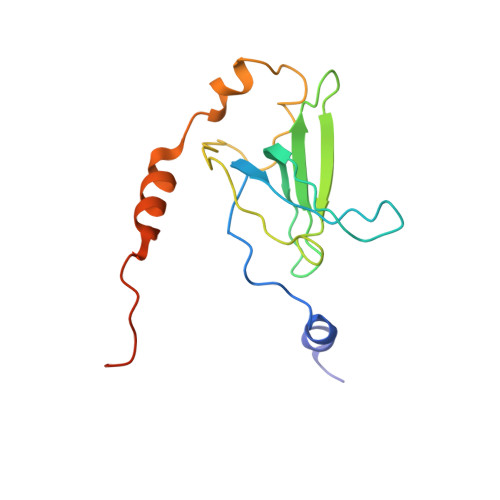
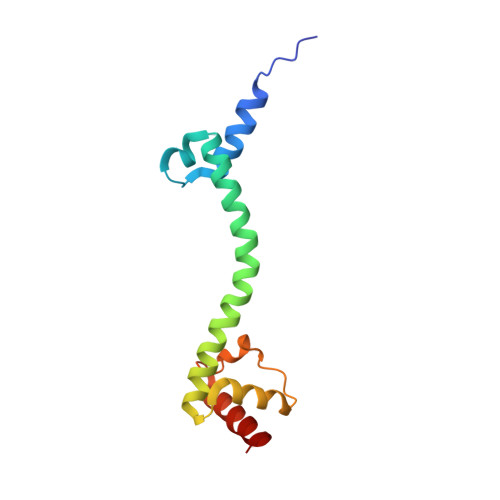
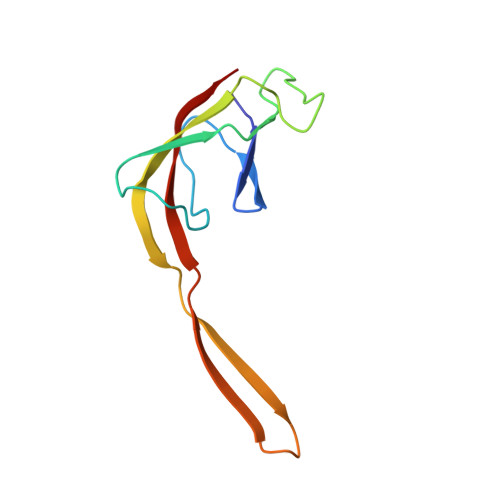
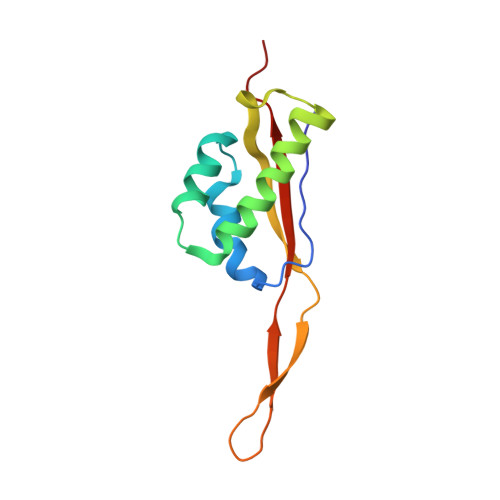
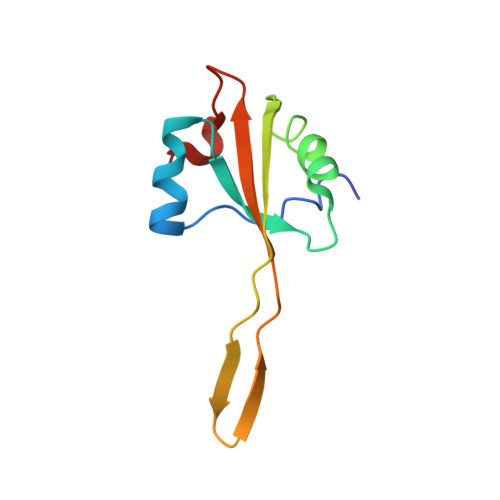
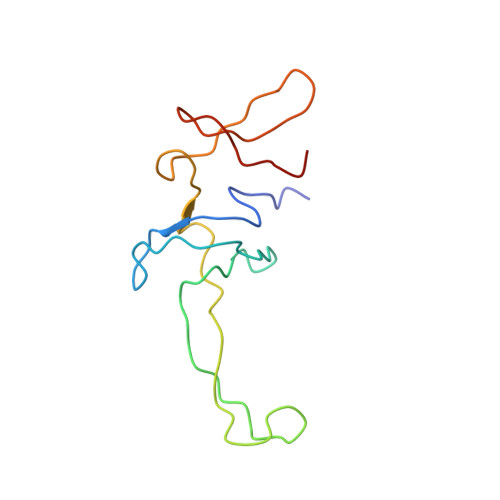
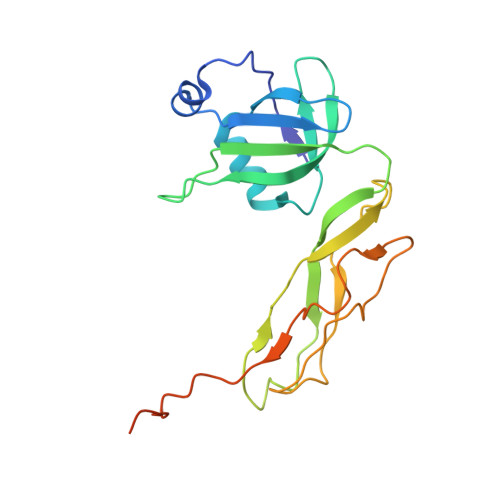
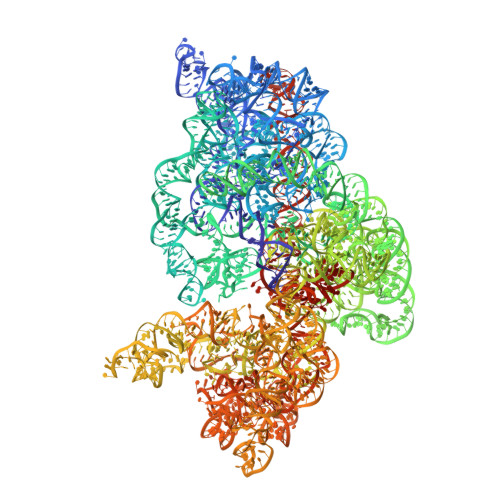
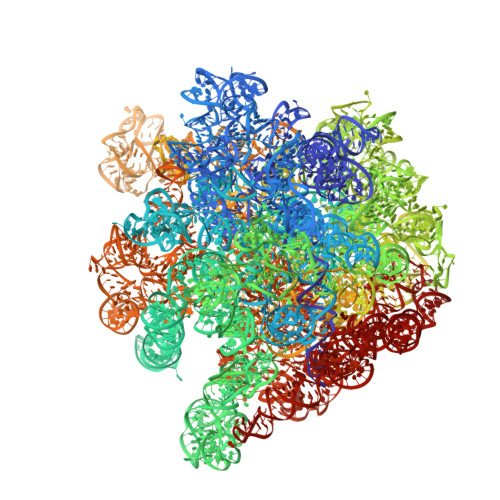
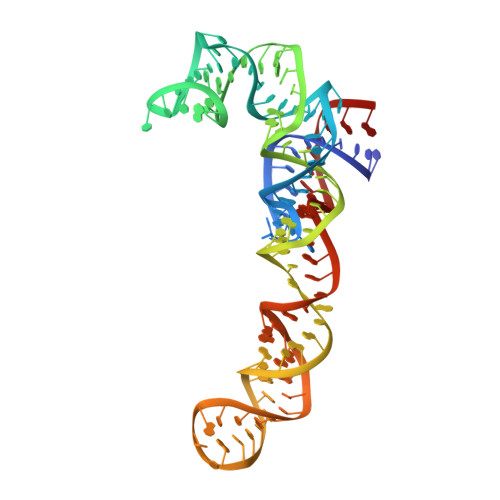
Crystal structure of 70S ribosome with both cognate tRNAs in the E and P sites representing an authentic elongation complex.
Feng, S., Chen, Y., Gao, Y.G.(2013) PLoS One 8: e58829-e58829
- PubMed: 23527033 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0058829
- Primary Citation Related Structures:
4V8U - PubMed Abstract:
During the translation cycle, a cognate deacylated tRNA can only move together with the codon into the E site. We here present the first structure of a cognate tRNA bound to the ribosomal E site resulting from translocation by EF-G, in which an entire L1 stalk (L1 protein and L1 rRNA) interacts with E-site tRNA (E-tRNA), representing an authentic ribosome elongation complex. Our results revealed that the Watson-Crick base pairing is formed at the first and second codon-anticodon positions in the E site in the ribosome elongation complex, whereas the codon-anticodon interaction in the third position is indirect. Analysis of the observed conformations of mRNA and E-tRNA suggests that the ribosome intrinsically has the potential to form codon-anticodon interaction in the E site, independently of the mRNA configuration. We also present a detailed description of the biologically relevant position of the entire L1 stalk and its interacting cognate E-tRNA, which provides a better understanding of the structural basis for translation elongation. Furthermore, to gain insight into translocation, we report the positioning of protein L6 contacting EF-G, as well as the conformational change of the C-terminal tail of protein S13 in the decoding center.
- School of Biological Science, Nanyang Technological University, Singapore.
Organizational Affiliation: