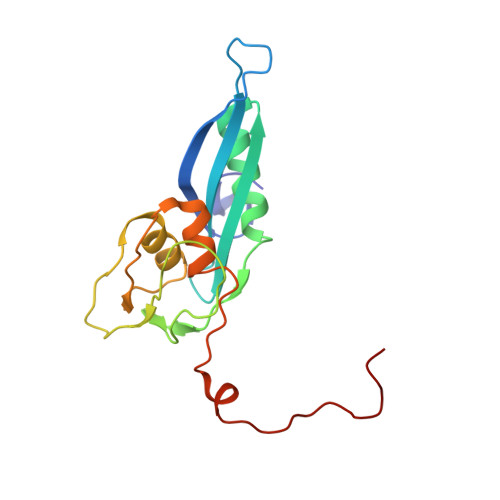
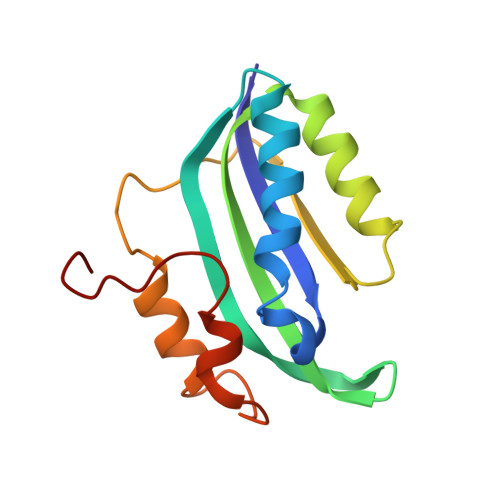
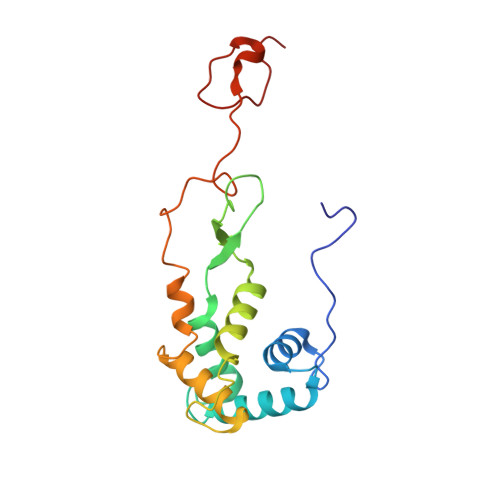
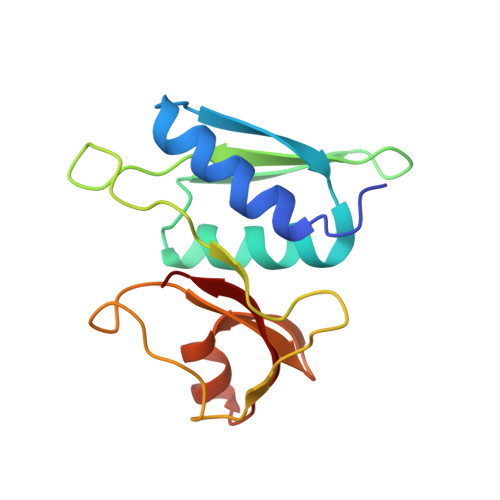
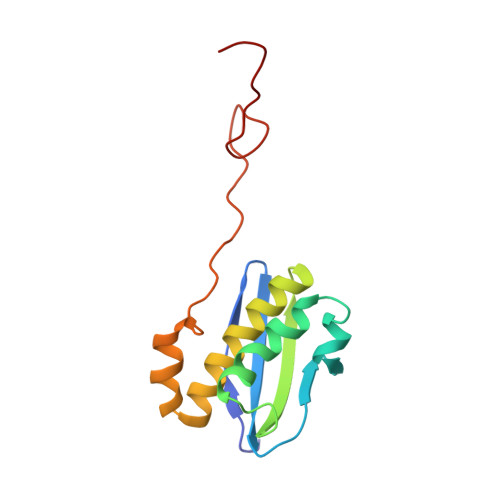
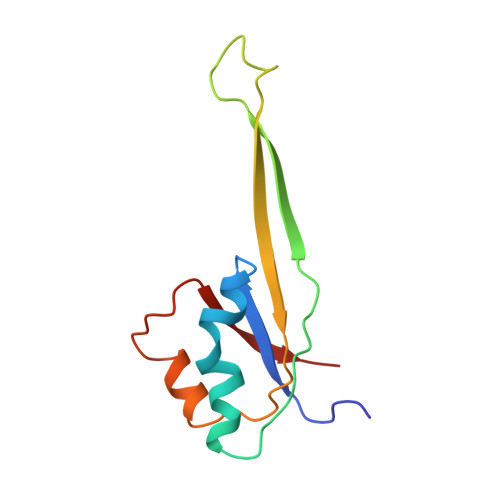
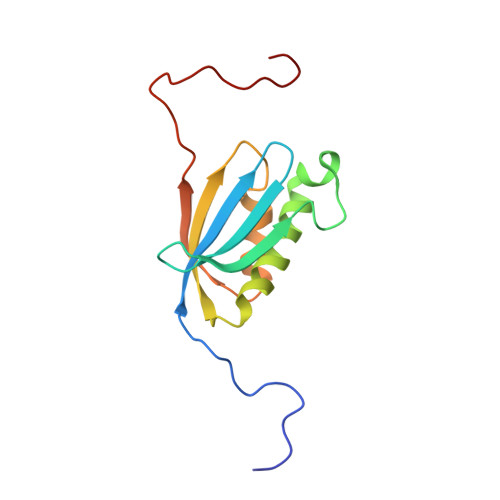
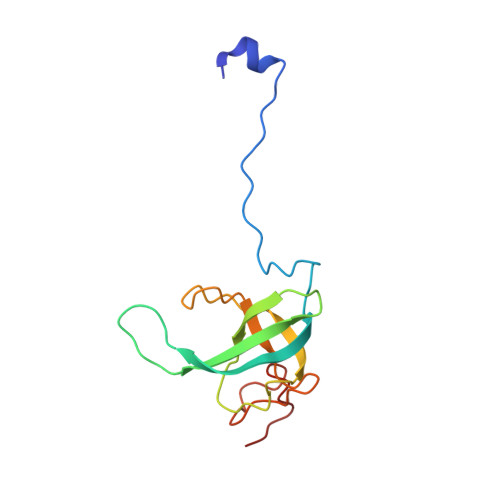
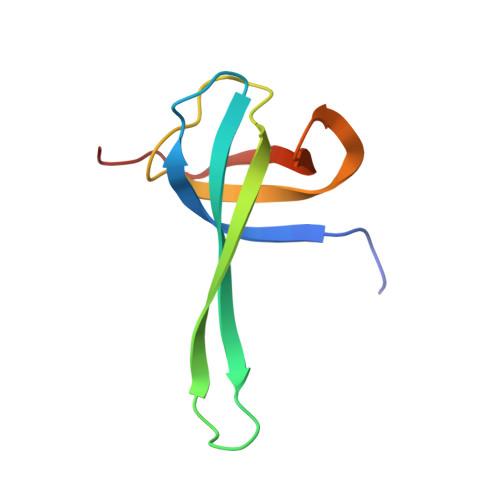
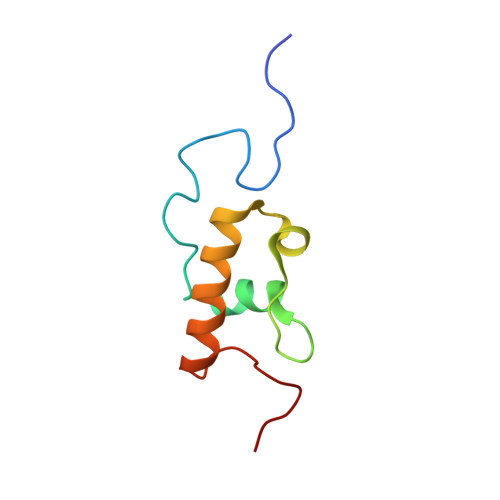
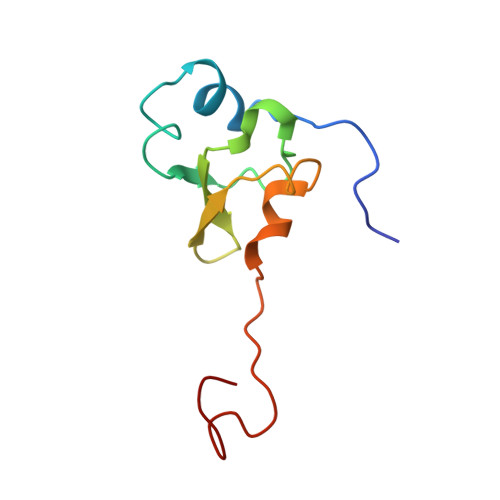
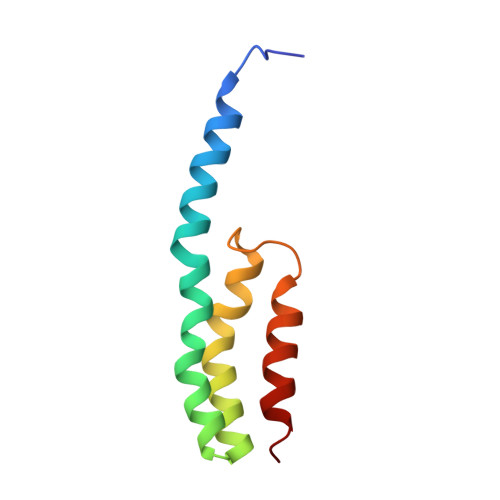
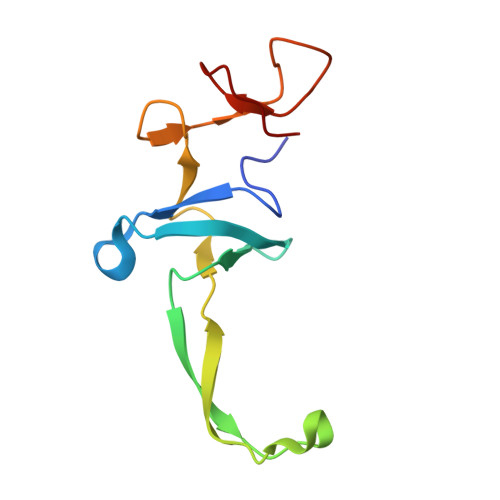
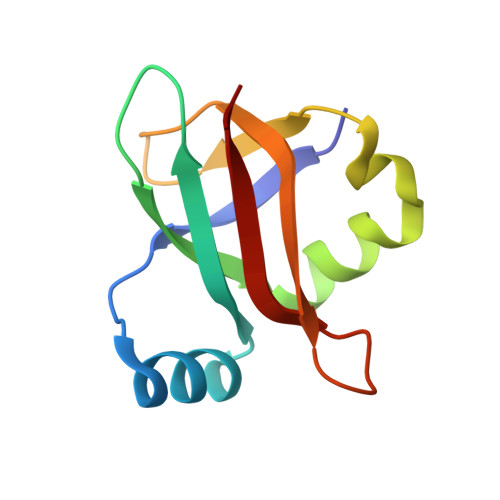
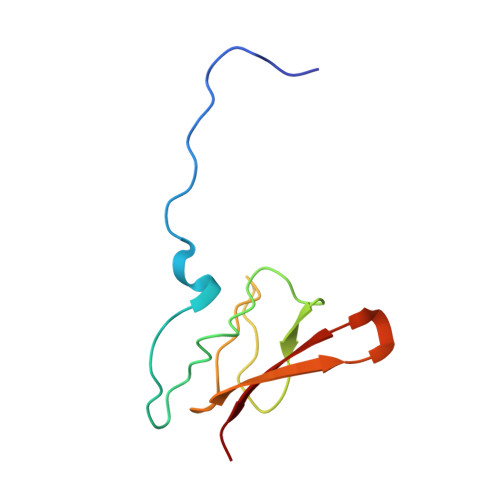
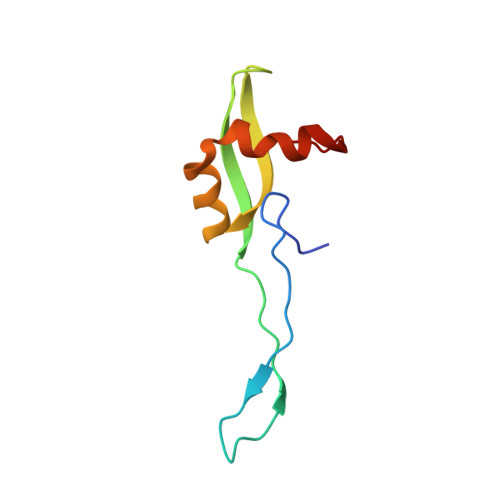
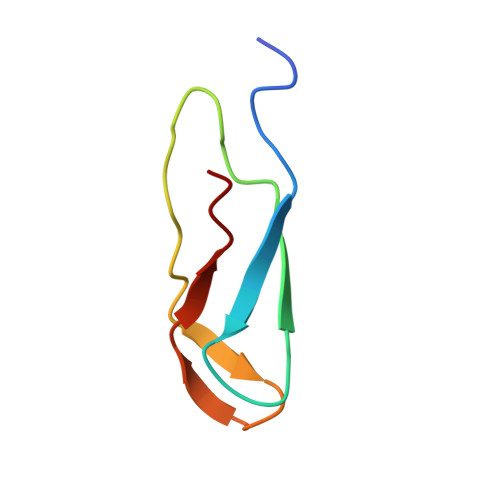
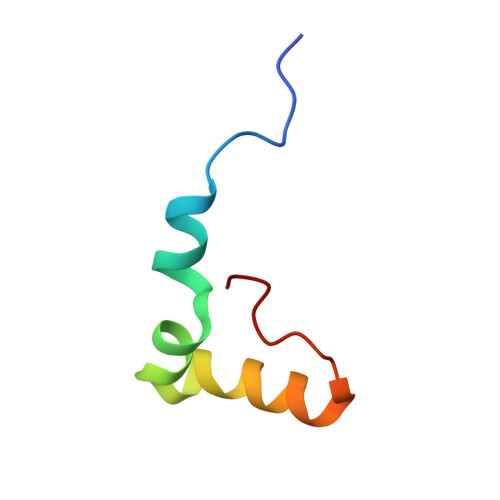
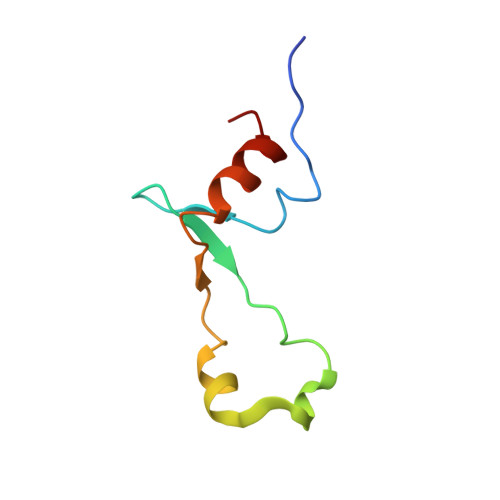
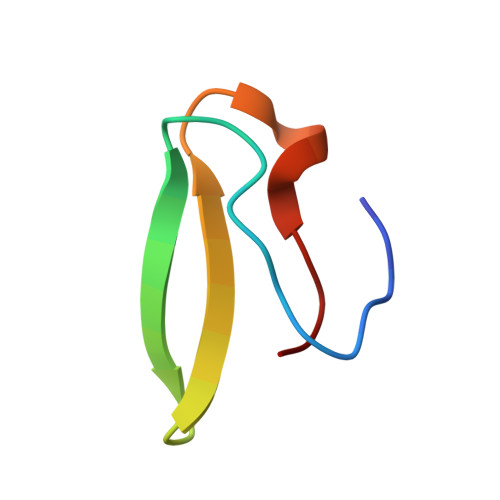
Structural insights into cognate versus near-cognate discrimination during decoding.
Agirrezabala, X., Schreiner, E., Trabuco, L.G., Lei, J., Ortiz-Meoz, R.F., Schulten, K., Green, R., Frank, J.(2011) EMBO J 30: 1497-1507
- PubMed: 21378755 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2011.58
- Primary Citation Related Structures:
4V6K, 4V6L - PubMed Abstract:
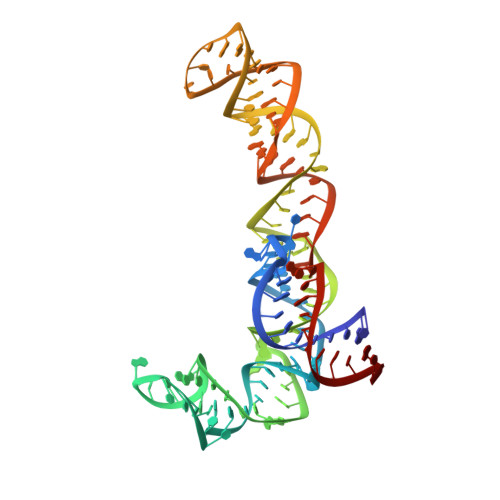
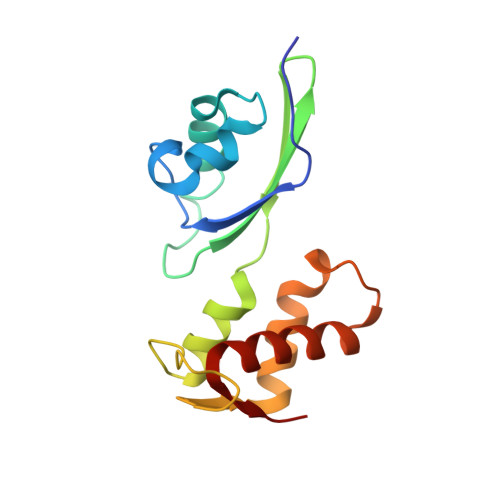
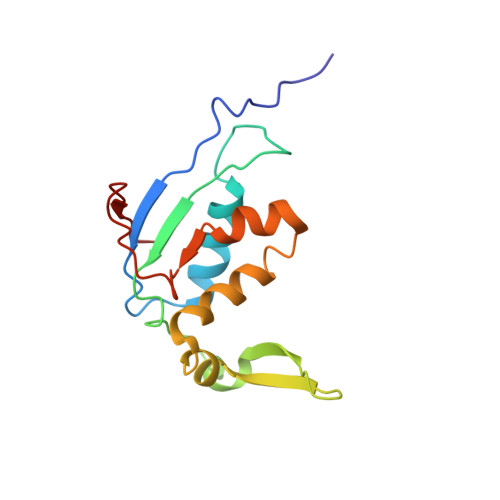
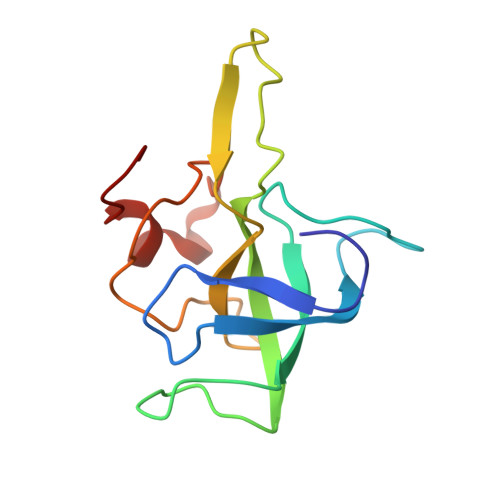
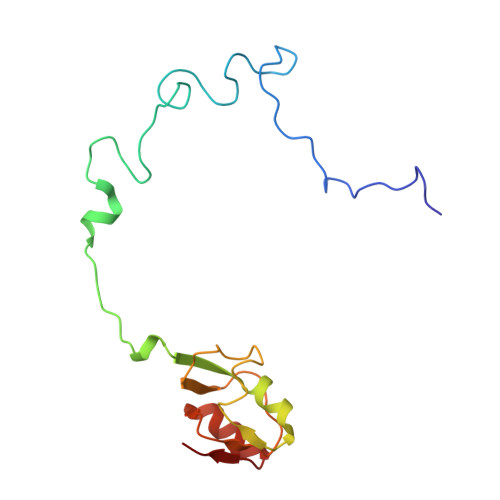
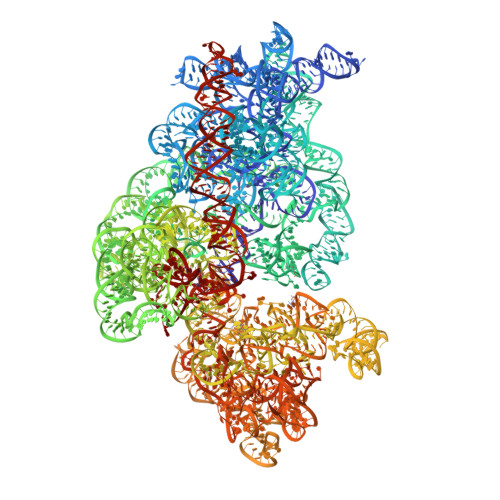
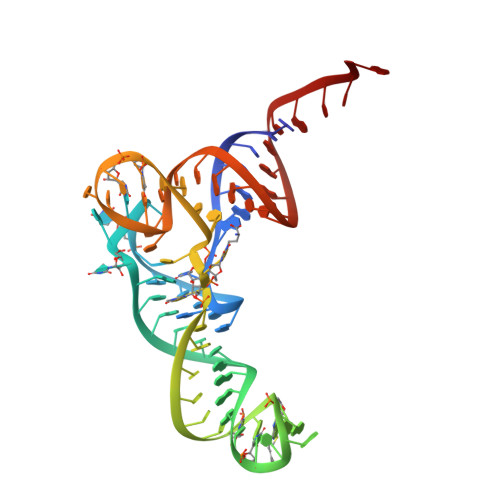
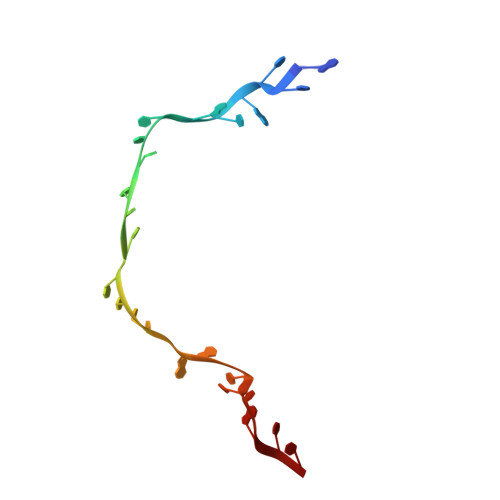
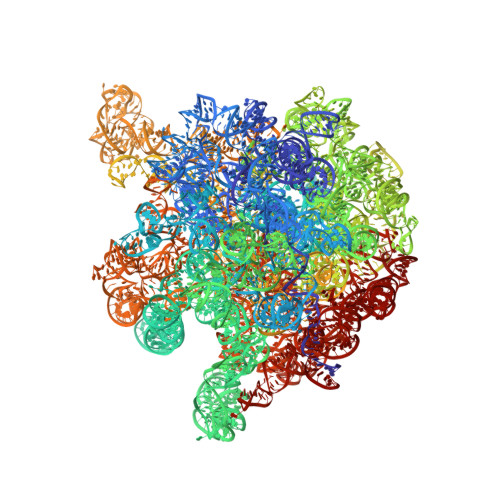
The structural basis of the tRNA selection process is investigated by cryo-electron microscopy of ribosomes programmed with UGA codons and incubated with ternary complex (TC) containing the near-cognate Trp-tRNA(Trp) in the presence of kirromycin. Going through more than 350 000 images and employing image classification procedures, we find ∼8% in which the TC is bound to the ribosome. The reconstructed 3D map provides a means to characterize the arrangement of the near-cognate aa-tRNA with respect to elongation factor Tu (EF-Tu) and the ribosome, as well as the domain movements of the ribosome. One of the interesting findings is that near-cognate tRNA's acceptor stem region is flexible and CCA end becomes disordered. The data bring direct structural insights into the induced-fit mechanism of decoding by the ribosome, as the analysis of the interactions between small and large ribosomal subunit, aa-tRNA and EF-Tu and comparison with the cognate case (UGG codon) offers clues on how the conformational signals conveyed to the GTPase differ in the two cases.
- Structural Biology Unit, CIC-bioGUNE, Derio, Basque Country, Spain.
Organizational Affiliation: