The Toolbox of Auricularia Auricula-Judae Dye-Decolorizing Peroxidase - Identification of Three New Potential Substrate-Interaction Sites.
Strittmatter, E., Serrer, K., Liers, C., Ullrich, R., Hofrichter, M., Piontek, K., Schleicher, E., Plattner, D.A.(2015) Arch Biochem Biophys 574: 75
- PubMed: 25542606 Search on PubMed
- DOI: https://doi.org/10.1016/j.abb.2014.12.016
- Primary Citation Related Structures:
4UZI - PubMed Abstract:
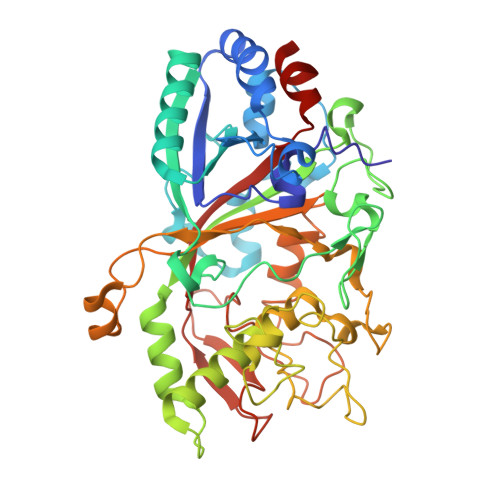
Dye-decolorizing peroxidases (DyPs) such as AauDyPI from the fungus Auricularia auricula-judae are able to oxidize substrates of different kinds and sizes. A crystal structure of an AauDyPI-imidazole complex gives insight into the binding patterns of organic molecules within the heme cavity of a DyP. Several small N-containing heterocyclic aromatics are shown to bind in the AauDyPI heme cavity, hinting to susceptibility of DyPs to azole-based inhibitors similar to cytochromes P450. Imidazole is confirmed as a competitive inhibitor with regard to peroxide binding. In contrast, bulky substrates such as anthraquinone dyes are converted at the enzyme surface. In the crystal structure a substrate analog, 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES), binds to a tyrosine-rich hollow harboring Y25, Y147, and Y337. Spin trapping with a nitric oxide donor uncovers Y229 as an additional tyrosine-based radical center in AauDyPI. Multi-frequency EPR spectroscopy further reveals the presence of at least one intermediate tryptophanyl radical center in activated AauDyPI with W377 as the most likely candidate.
- Institute of Organic Chemistry, University of Freiburg, Albertstrasse 21, 79104 Freiburg, Germany.
Organizational Affiliation: