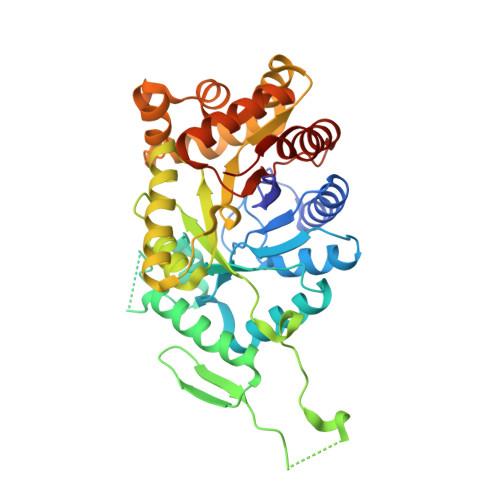
The Oxygenating Constituent of 3,6-Diketocamphane Monooxygenase from the Cam Plasmid of Pseudomonas Putida: The First Crystal Structure of a Type II Baeyer-Villiger Monooxygenase.
Isupov, M.N., Schroder, E., Gibson, R.P., Beecher, J., Donadio, G., Saneei, V., Dcunha, S.A., Mcghie, E.J., Sayer, C., Davenport, C.F., Lau, P.C., Hasegawa, Y., Iwaki, H., Kadow, M., Balke, K., Bornscheuer, U.T., Bourenkov, G., Littlechild, J.A.(2015) Acta Crystallogr D Biol Crystallogr 71: 2344
- PubMed: 26527149 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004715017939
- Primary Citation Related Structures:
4UWM, 5AEC - PubMed Abstract:
The three-dimensional structures of the native enzyme and the FMN complex of the overexpressed form of the oxygenating component of the type II Baeyer-Villiger 3,6-diketocamphane monooxygenase have been determined to 1.9 Å resolution. The structure of this dimeric FMN-dependent enzyme, which is encoded on the large CAM plasmid of Pseudomonas putida, has been solved by a combination of multiple anomalous dispersion from a bromine crystal soak and molecular replacement using a bacterial luciferase model. The orientation of the isoalloxazine ring of the FMN cofactor in the active site of this TIM-barrel fold enzyme differs significantly from that previously observed in enzymes of the bacterial luciferase-like superfamily. The Ala77 residue is in a cis conformation and forms a β-bulge at the C-terminus of β-strand 3, which is a feature observed in many proteins of this superfamily.
- The Henry Wellcome Building for Biocatalysis, Biosciences, College of Life and Environmental Sciences, University of Exeter, Stocker Road, Exeter EX4 4QD, England.
Organizational Affiliation: