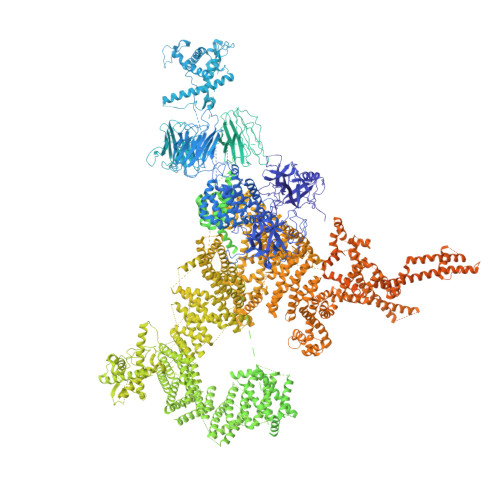
Architecture and Conformational Switch Mechanism of the Ryanodine Receptor.
Efremov, R.G., Leitner, A., Aebersold, R., Raunser, S.(2015) Nature 517: 39
- PubMed: 25470059 Search on PubMed
- DOI: https://doi.org/10.1038/nature13916
- Primary Citation Related Structures:
4UWA, 4UWE - PubMed Abstract:
Muscle contraction is initiated by the release of calcium (Ca(2+)) from the sarcoplasmic reticulum into the cytoplasm of myocytes through ryanodine receptors (RyRs). RyRs are homotetrameric channels with a molecular mass of more than 2.2 megadaltons that are regulated by several factors, including ions, small molecules and proteins. Numerous mutations in RyRs have been associated with human diseases. The molecular mechanism underlying the complex regulation of RyRs is poorly understood. Using electron cryomicroscopy, here we determine the architecture of rabbit RyR1 at a resolution of 6.1 Å. We show that the cytoplasmic moiety of RyR1 contains two large α-solenoid domains and several smaller domains, with folds suggestive of participation in protein-protein interactions. The transmembrane domain represents a chimaera of voltage-gated sodium and pH-activated ion channels. We identify the calcium-binding EF-hand domain and show that it functions as a conformational switch allosterically gating the channel.
- 1] Department of Structural Biochemistry, Max Planck Institute of Molecular Physiology, 44227 Dortmund, Germany [2] Structural Biology Research Center, Vlaams Instituut voor Biotechnologie (VIB), 1050 Brussels, Belgium [3] Structural Biology Brussels, Vrije Universiteit Brussel (VUB), 1050 Brussels, Belgium.
Organizational Affiliation: