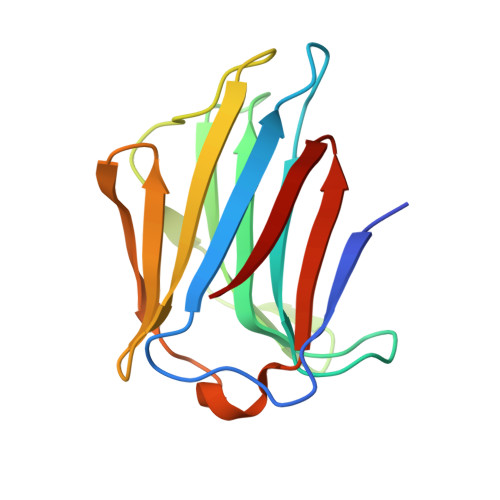
Structural Basis of Multivalent Galactose-Based Dendrimer Recognition by Human Galectin-7.
Ramaswamy, S., Haj Sleiman, M., Masuyer, G., Arbez-Gindre, C., Micha-Screttas, M., Calogeropoulou, T., Steele, B.R., Acharya, K.R.(2015) FEBS J 282: 372
- PubMed: 25367374 Search on PubMed
- DOI: https://doi.org/10.1111/febs.13140
- Primary Citation Related Structures:
4UW3, 4UW4, 4UW5, 4UW6 - PubMed Abstract:
Galectins are evolutionarily conserved and ubiquitously present animal lectins with a high affinity for β-galactose-containing oligosaccharides. To date, 15 mammalian galectins have been identified. Their involvement in cell-cell and cell-matrix interactions has highlighted their importance in signal transduction and other intracellular processes. Human galectin-7 (hGal-7) is a 15 kDa proto type galectin that forms a dimer in solution and its involvement in the stimulation and development of tumour growth has been reported. Previously, we reported the crystal structure of hGal-7 and its complex with galactose and lactose which provided insight into its molecular recognition and detailed interactions. Here, we present newly obtained high-resolution structural data on carbohydrate-based dendrons in complex with hGal-7. Our crystallographic data reveal how multivalent ligands interact with and form cross-links with these galectin molecules. Understanding how these dendrimeric compounds interact with hGal-7 would help in the design of new tools to investigate the recognition of carbohydrates by lectins.
- Department of Biology and Biochemistry, University of Bath, UK.
Organizational Affiliation: