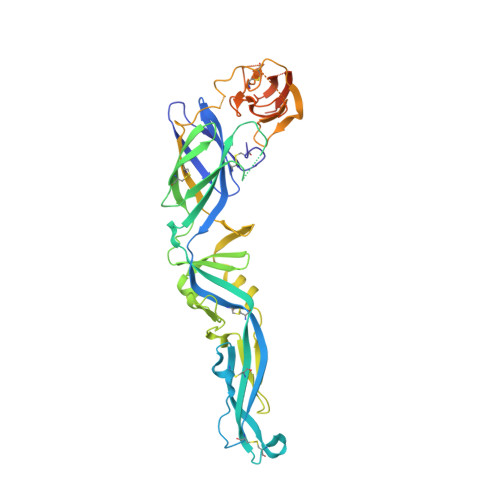
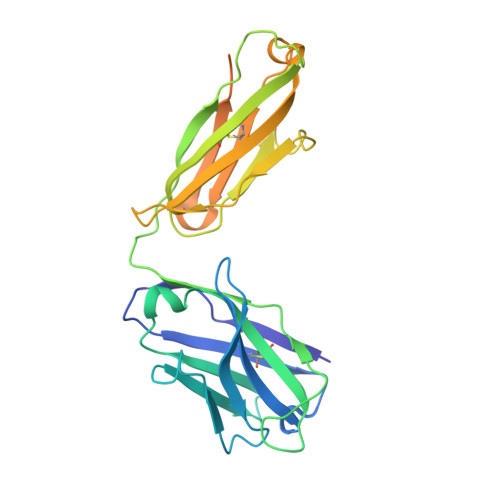
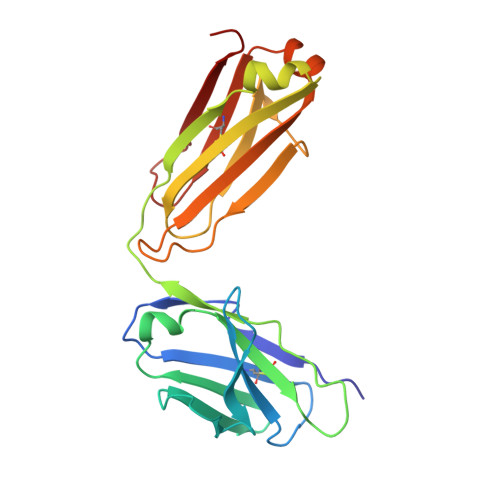
Recognition Determinants of Broadly Neutralizing Human Antibodies Against Dengue Viruses.
Rouvinski, A., Guardado-Calvo, P., Barba-Spaeth, G., Duquerroy, S., Vaney, M., Kikuti, C.M., Sanchez, M.E.N., Dejnirattisai, W., Wongwiwat, W., Haouz, A., Girard-Blanc, C., Petres, S., Shepard, W.E., Despres, P., Arenzana-Seisdedos, F., Dussart, P., Mongkolsapaya, J., Screaton, G.R., Rey, F.A.(2015) Nature 520: 109
- PubMed: 25581790 Search on PubMed
- DOI: https://doi.org/10.1038/nature14130
- Primary Citation Related Structures:
4UT6, 4UT7, 4UT9, 4UTA, 4UTB, 4UTC - PubMed Abstract:
Dengue disease is caused by four different flavivirus serotypes, which infect 390 million people yearly with 25% symptomatic cases and for which no licensed vaccine is available. Recent phase III vaccine trials showed partial protection, and in particular no protection for dengue virus serotype 2 (refs 3, 4). Structural studies so far have characterized only epitopes recognized by serotype-specific human antibodies. We recently isolated human antibodies potently neutralizing all four dengue virus serotypes. Here we describe the X-ray structures of four of these broadly neutralizing antibodies in complex with the envelope glycoprotein E from dengue virus serotype 2, revealing that the recognition determinants are at a serotype-invariant site at the E-dimer interface, including the exposed main chain of the E fusion loop and the two conserved glycan chains. This 'E-dimer-dependent epitope' is also the binding site for the viral glycoprotein prM during virus maturation in the secretory pathway of the infected cell, explaining its conservation across serotypes and highlighting an Achilles' heel of the virus with respect to antibody neutralization. These findings will be instrumental for devising novel immunogens to protect simultaneously against all four serotypes of dengue virus.
- 1] Institut Pasteur, Département de Virologie, Unité de Virologie Structurale, 75724 Paris Cedex 15, France [2] CNRS UMR 3569 Virologie, 75724 Paris Cedex 15, France.
Organizational Affiliation: