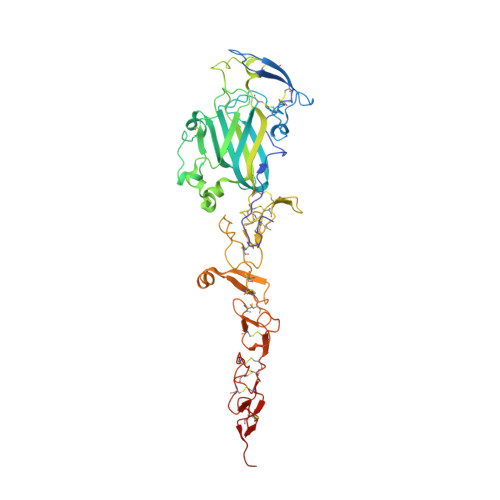
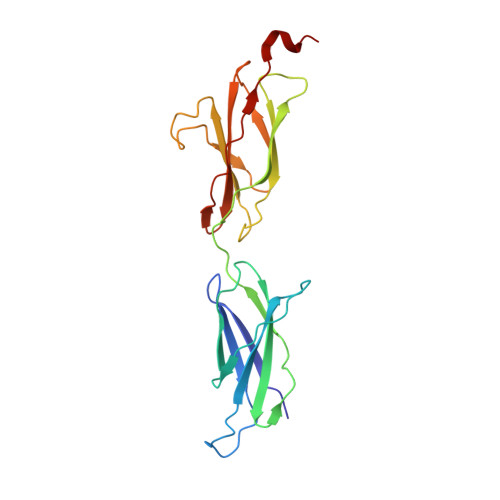
The Crystal Structure of Netrin-1 in Complex with Dcc Reveals the Bi-Functionality of Netrin-1 as a Guidance Cue
Finci, L.I., Krueger, N., Sun, X., Zhang, J., Chegkazi, M., Wu, Y., Schenk, G., Mertens, H.D.T., Svergun, D.I., Zhang, Y., Wang, J.-H., Meijers, R.(2014) Neuron 83: 839
- PubMed: 25123307 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.neuron.2014.07.010
- Primary Citation Related Structures:
4URT - PubMed Abstract:
Netrin-1 is a guidance cue that can trigger either attraction or repulsion effects on migrating axons of neurons, depending on the repertoire of receptors available on the growth cone. How a single chemotropic molecule can act in such contradictory ways has long been a puzzle at the molecular level. Here we present the crystal structure of netrin-1 in complex with the Deleted in Colorectal Cancer (DCC) receptor. We show that one netrin-1 molecule can simultaneously bind to two DCC molecules through a DCC-specific site and through a unique generic receptor binding site, where sulfate ions staple together positively charged patches on both DCC and netrin-1. Furthermore, we demonstrate that UNC5A can replace DCC on the generic receptor binding site to switch the response from attraction to repulsion. We propose that the modularity of binding allows for the association of other netrin receptors at the generic binding site, eliciting alternative turning responses.
- State Key Laboratory of Biomembrane and Membrane Biotechnology, College of Life Sciences, Peking University, Beijing, 100871, China.
Organizational Affiliation: