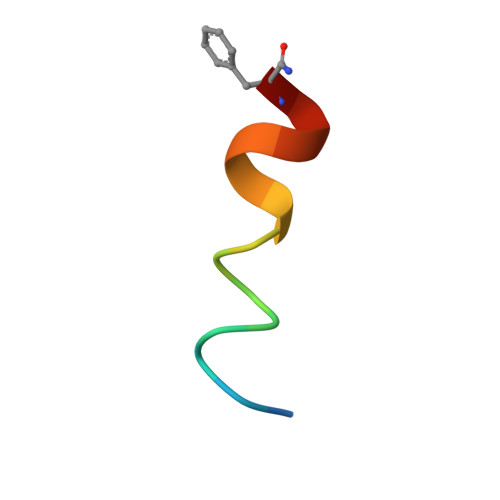
Benzene Probes in Molecular Dynamics Simulations Reveal Novel Binding Sites for Ligand Design.
Tan, Y.S., Reeks, J., Brown, C.J., Thean, D., Ferrer Gago, F.J., Yuen, T.Y., Goh, E.T., Lee, X.E., Jennings, C.E., Joseph, T.L., Lakshminarayanan, R., Lane, D.P., Noble, M.E., Verma, C.S.(2016) J Phys Chem Lett 7: 3452-3457
- PubMed: 27532490 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jpclett.6b01525
- Primary Citation Related Structures:
4UD7, 4UE1 - PubMed Abstract:
Protein flexibility poses a major challenge in binding site identification. Several computational pocket detection methods that utilize small-molecule probes in molecular dynamics (MD) simulations have been developed to address this issue. Although they have proven hugely successful at reproducing experimental structural data, their ability to predict new binding sites that are yet to be identified and characterized has not been demonstrated. Here, we report the use of benzenes as probe molecules in ligand-mapping MD (LMMD) simulations to predict the existence of two novel binding sites on the surface of the oncoprotein MDM2. One of them was serendipitously confirmed by biophysical assays and X-ray crystallography to be important for the binding of a new family of hydrocarbon stapled peptides that were specifically designed to target the other putative site. These results highlight the predictive power of LMMD and suggest that predictions derived from LMMD simulations can serve as a reliable basis for the identification of novel ligand binding sites in structure-based drug design.
- Bioinformatics Institute, Agency for Science, Technology and Research (A*STAR) , 30 Biopolis Street, #07-01 Matrix, Singapore 138671.
Organizational Affiliation: