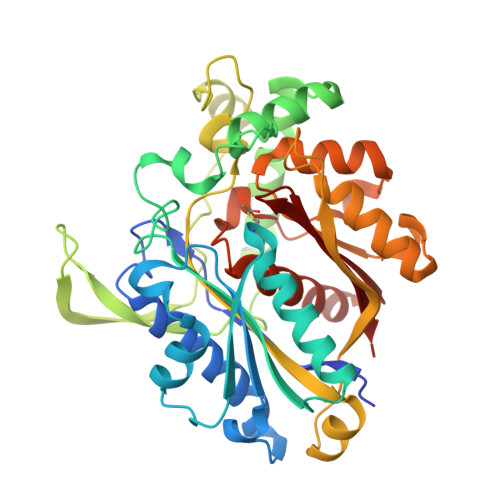
FadA5 a Thiolase from Mycobacterium tuberculosis: A Steroid-Binding Pocket Reveals the Potential for Drug Development against Tuberculosis.
Schaefer, C.M., Lu, R., Nesbitt, N.M., Schiebel, J., Sampson, N.S., Kisker, C.(2015) Structure 23: 21-33
- PubMed: 25482540 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2014.10.010
- Primary Citation Related Structures:
4UBT, 4UBU, 4UBV, 4UBW - PubMed Abstract:
With the exception of HIV, tuberculosis (TB) is the leading cause of mortality among infectious diseases. The urgent need to develop new antitubercular drugs is apparent due to the increasing number of drug-resistant Mycobacterium tuberculosis (Mtb) strains. Proteins involved in cholesterol import and metabolism have recently been discovered as potent targets against TB. FadA5, a thiolase from Mtb, is catalyzing the last step of the β-oxidation reaction of the cholesterol side-chain degradation under release of critical metabolites and was shown to be of importance during the chronic stage of TB infections. To gain structural and mechanistic insight on FadA5, we characterized the enzyme in different stages of the cleavage reaction and with a steroid bound to the binding pocket. Structural comparisons to human thiolases revealed that it should be possible to target FadA5 specifically, and the steroid-bound structure provides a solid basis for the development of inhibitors against FadA5.
- Rudolf Virchow Center for Experimental Biomedicine, Institute for Structural Biology, University of Würzburg, Josef-Schneider-Strasse 2, 97080 Würzburg, Germany.
Organizational Affiliation: