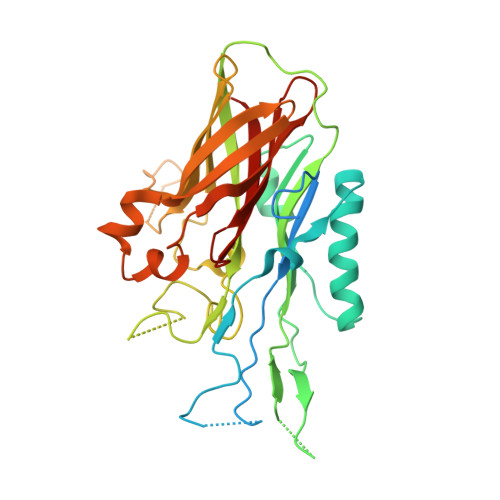
The structure of lactoferrin-binding protein B from Neisseria meningitidis suggests roles in iron acquisition and neutralization of host defences.
Brooks, C.L., Arutyunova, E., Lemieux, M.J.(2014) Acta Crystallogr F Struct Biol Commun 70: 1312-1317
- PubMed: 25286931
- DOI: https://doi.org/10.1107/S2053230X14019372
- Primary Citation Related Structures:
4U9C - PubMed Abstract:
Pathogens have evolved a range of mechanisms to acquire iron from the host during infection. Several Gram-negative pathogens including members of the genera Neisseria and Moraxella have evolved two-component systems that can extract iron from the host glycoproteins lactoferrin and transferrin. The homologous iron-transport systems consist of a membrane-bound transporter and an accessory lipoprotein. While the mechanism behind iron acquisition from transferrin is well understood, relatively little is known regarding how iron is extracted from lactoferrin. Here, the crystal structure of the N-terminal domain (N-lobe) of the accessory lipoprotein lactoferrin-binding protein B (LbpB) from the pathogen Neisseria meningitidis is reported. The structure is highly homologous to the previously determined structures of the accessory lipoprotein transferrin-binding protein B (TbpB) and LbpB from the bovine pathogen Moraxella bovis. Docking the LbpB structure with lactoferrin reveals extensive binding interactions with the N1 subdomain of lactoferrin. The nature of the interaction precludes apolactoferrin from binding LbpB, ensuring the specificity of iron-loaded lactoferrin. The specificity of LbpB safeguards proper delivery of iron-bound lactoferrin to the transporter lactoferrin-binding protein A (LbpA). The structure also reveals a possible secondary role for LbpB in protecting the bacteria from host defences. Following proteolytic digestion of lactoferrin, a cationic peptide derived from the N-terminus is released. This peptide, called lactoferricin, exhibits potent antimicrobial effects. The docked model of LbpB with lactoferrin reveals that LbpB interacts extensively with the N-terminal lactoferricin region. This may provide a venue for preventing the production of the peptide by proteolysis, or directly sequestering the peptide, protecting the bacteria from the toxic effects of lactoferricin.
- Department of Chemistry, California State University Fresno, Fresno, CA 93710, USA.
Organizational Affiliation: